Difference between revisions of "20.109(F08): Mod 2 Day 3 Colony PCR and journal article discussion"
From Course Wiki
(→Reagents list) |
MAXINE JONAS (Talk | contribs) m (22 revisions: Transfer 20.109(F08) to HostGator) |
||
| (18 intermediate revisions by 2 users not shown) | |||
| Line 2: | Line 2: | ||
==Introduction== | ==Introduction== | ||
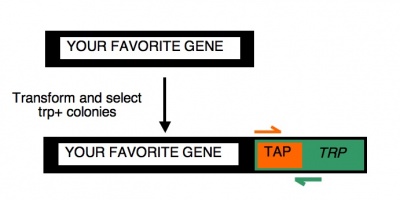
| − | To begin | + | To begin today���s experiment, you will ���pop��� some trp+ yeast from your transformation plates and then amplify the relevant portion of the released genomic DNA. The primers you will use are expected to give a ~554 base pair product only if the TAP-TRP tag is present. [[Image:CheckingTAP.jpg|400px|thumb|right| PCR primers to check candidates]] This is accomplished by using a forward primer that anneals to the very 5'-end of the TAP gene you've tried to add to your gene of interest and a reverse primer that anneals to a very 3'-end of the TRP gene that was part of the TAP tag and used for selection. There are two important caveats about today's experiment. First, if the TAP tag is fused elsewhere in the genome (giving rise to the trp+ phenotype) but the SAGA- or SAGA-controlled gene remains intact, then you may still see a product from this reaction. Thus we'll have to follow up in other ways to confirm that the gene of interest has, in fact, been modified. Second, you should be aware that a negative result from these reactions (i.e. no PCR product) can just as easily be explained as a failed PCR (bad primers, dead enzyme, wrong reaction conditions, etc) as a failed tagging transformation. So don't let a negative result today fool you and lead you to toss out perfectly correct samples. Only the positive result is meaningful in this experiment and we will not know the result until you run the agarose gel next time. So, we will remain optimistic and set up overnight cultures of both the candidates you are examining today. In this way you will have cells to examine next time, when you will check them for the TAP-tag by Western blot and when you will isolate total RNA from them for microarray analysis. |
| − | + | ||
==Protocols== | ==Protocols== | ||
===Part 1: Colony PCR=== | ===Part 1: Colony PCR=== | ||
| − | #Begin by counting the colonies that arose from the transformation experiment you performed last time. Choose | + | #Begin by counting the colonies that arose from the transformation experiment you performed last time. Choose two candidates (when possible) to pursue, circling the colonies on the back of the petri dish and clearly labeling them "A," and "B."If you do not have enough colonies, don't despair. You can get some additional ones from another group or from the teaching faculty. Note, however, that by doing so, you may be switching the gene you'll study for the rest of the module, and you will have to rethink the work you have planned so far. And if applicable, it will be worth noting the "negative" result in the research article you write, since it may indicate that despite the gene being deemed non-essential, cells may fare less well than wild-type cells when the gene is modified and so modified forms may be difficult to isolate. |
| − | #You will use the microwave to release the DNA from the yeast. On the tip of a sterile toothpick, pick-up 1/2 of the colony that is candidate "A" and swirl the cells in 20 μl of sterile water in an eppendorf tube. Be sure to label the tube so you know it belongs to your group and which candidate it contains. Repeat with the second | + | #You will use the microwave to release the DNA from the yeast. On the tip of a sterile toothpick, pick-up 1/2 of the colony that is candidate "A" and swirl the cells in 20 μl of sterile water in an eppendorf tube. Be sure to label the tube so you know it belongs to your group and which candidate it contains. Repeat with the second candidate you've selected as well as with the trp+ version of the parental strain, NY420 and a positive control strain for this part of the experiment, either NY412 (same as NY411 but SPT3-TAP tagged) or NY413 (same as NY411 but SPT7-TAP tagged). The teaching faculty will have the control strains streaked out on petri dishes. You should mix 1/2 of a colony from these plate with 20 μl of sterile water as you did for your candidate colonies. |
| − | + | #Close the caps to the four tubes and microwave them in an eppendorf rack for 15 seconds. | |
| − | #Close the caps to the tubes and microwave them in an eppendorf rack for 15 seconds. | + | |
#Move 2.5 μl of the microwaved mixes, yeast debris and all, into a 200 μl PCR tube. Again label your tubes well (write small!). | #Move 2.5 μl of the microwaved mixes, yeast debris and all, into a 200 μl PCR tube. Again label your tubes well (write small!). | ||
| − | #In a new eppendorf tube (normal size this time), prepare a PCR cocktail enough for 5 reactions. Each reaction will include: | + | #Prepare your PCR cocktails. These are best performed with a reaction cocktail that contains the reagents common to all 4 reactions. In a new eppendorf tube (normal size this time), prepare a PCR cocktail enough for 5 reactions since you'll want a little extra volume to account for pipetting error. Each reaction will include: |
#*10 μl 2.5X PCR mastermix | #*10 μl 2.5X PCR mastermix | ||
| − | #*0.5 μl forward primer specific to | + | #*0.5 μl universal forward primer NO224, specific to the TAP portion of your tag |
| − | #*0.5 μl reverse primer specific | + | #*0.5 μl universal reverse primer NO225, specific to the TRP portion of your tag |
#*water to a final volume of 22.5 μl. | #*water to a final volume of 22.5 μl. | ||
| − | #Add 22.5 μl of PCR cocktail to each PCR tube with yeast and leave the tubes | + | #Add 22.5 μl of PCR cocktail to each PCR tube with yeast and leave the tubes chilling until everyone is ready. |
#Cycle the reactions as: | #Cycle the reactions as: | ||
#*95° 4 minutes | #*95° 4 minutes | ||
#*95° 1 minute | #*95° 1 minute | ||
#*52° 1 minute | #*52° 1 minute | ||
| − | #*72° 2 | + | #*72° 2 minutes |
#*repeat steps 2-4 35 times | #*repeat steps 2-4 35 times | ||
#*72° 10 minutes | #*72° 10 minutes | ||
| Line 28: | Line 26: | ||
===Part 2: Overnight cultures=== | ===Part 2: Overnight cultures=== | ||
| − | #Using sterile technique, aliquot | + | #Using sterile technique, aliquot 3 ml of YPD into four sterile test tubes. |
| − | #Label the caps with your team color, and either "parent" " | + | #Label the caps with your team color, and either "parent" "+control" "A" or "B." |
#Use sterile dowels to move the second 1/2 of each yeast colony from your transformation plates to the media. Swirl the dowel to remove the yeast from the stick and vortex the solution to fully resuspend them. | #Use sterile dowels to move the second 1/2 of each yeast colony from your transformation plates to the media. Swirl the dowel to remove the yeast from the stick and vortex the solution to fully resuspend them. | ||
| − | #Use a sterile dowel to innoculate the "parent" | + | #Use a sterile dowel to innoculate the "parent" and "+control" cultures, using the NY420 and NY412 or NY413 plates that the teaching faculty have prepared. |
| − | # | + | #Leave your tubes with the teaching faculty who will place them on the roller drum in the 30�� incubator. The cultures will grow for 24 hours and then be placed in the 4�� fridge until next lab. |
#When everyone is ready we will discuss the journal article that was assigned for today. | #When everyone is ready we will discuss the journal article that was assigned for today. | ||
| + | |||
===Part 3: Journal article discussion=== | ===Part 3: Journal article discussion=== | ||
| − | As part of your assignment for today, you have read the relevant article by [http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?db=pubmed&cmd=Retrieve&dopt=AbstractPlus&list_uids=15260971&query_hl=3&itool=pubmed_docsum Wu et al, published in Mol Cell in 2004]. There is also an associated review article [ | + | As part of your assignment for today, you have read the relevant article by [http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?db=pubmed&cmd=Retrieve&dopt=AbstractPlus&list_uids=15260971&query_hl=3&itool=pubmed_docsum Wu et al, published in Mol Cell in 2004]. There is also an associated review article [http://openwetware.org/wiki/PMID:_15653319 ] that was written to celebrate and highlight this important work on the structure of the SAGA complex. You and your partner will be randomly assigned a portion of the text to describe to the class but be prepared to engage in conversation about other parts of this paper even if you're not "assigned" to the section being discussed. |
DONE! | DONE! | ||
==For next time== | ==For next time== | ||
| − | # | + | #Write a paragraph to report on your transformation results. Eventually this homework assignment will be integrated into the results section for your [[20.109(F08): Protein engineering research article| research article]] that you'll hand in a few weeks from now. Report what you did and what you found here. Leave interpretation to the next FNT question. |
| + | #Write a sentence or two to interpret your transformation results. Eventually this information will be integrated into the discussion section of your [[20.109(F08): Protein engineering research article| research article]] that you'll hand in a few weeks from now. | ||
#You are not required to hand in a revised Materials and Methods section, but consider adding the experiment you performed today to the assignment from last time. | #You are not required to hand in a revised Materials and Methods section, but consider adding the experiment you performed today to the assignment from last time. | ||
| + | #Next lab may run long!!! Plan ahead and come prepared with your strategy for testing phenotypes of your strains. What kinds of media are you interested in testing? If you'd like media besides YPD, YPGAL, -trp and -his, you must let us know one day in advance of lab so we can prepare them for you. | ||
| + | #Sign up for one of the two Journal Club presentations days [[20.109(F08): Sign up for journal club| here.]] We must split the class evenly between the two available days. First come first served. You do not have to select a paper to present yet (unless you want to!). | ||
==Reagents list== | ==Reagents list== | ||
| + | *YPD | ||
| + | **10g yeast extract | ||
| + | **20g peptone | ||
| + | **20g glucose | ||
| + | **20g agar (for liquid media, leave agar out!) | ||
| + | **add 1L water, autoclave 20min, stir to cool | ||
| + | *PCR reaction mix | ||
| + | *Primers | ||
| + | **Universal forward primer | ||
| + | ***'''NO224''' TCC ATG GAA AAG AGA AGA TGG AAA AAG AAT TTC ATA GCC GTC TC | ||
| + | ***use with rev primer for checking TAP-TRP tag by PCR | ||
| + | ***44 mer | ||
| + | ***Tm = 63.6 | ||
| + | **Universal reverse primer | ||
| + | ***'''NO225''' TCA GGT TGA CTT CCC CGC GGA ATT CGC GTC TAC | ||
| + | ***use with fwd primer for checking TAP-TRP tag by PCR | ||
| + | ***expect length of TAP seq + TRP K. l seq = 554 bps (from 2147 to 2701 in pBS1479) | ||
| + | ***Tm = 68 | ||
| + | **Other positive control primers (not used today) | ||
| + | ***'''NO258''' for SPT3-TAP strain NY412, TTT AAA GGT GGT AGA CTC AGT TCT AAA CCA ATT ATC ATG tcc atg gaa aag aga aga tg | ||
| + | ***'''NO259''' for SPT7-TAP strain NY413, AAT AGT TCA TTT AGC TTG AGC CTT CCT CGC CTT AAT CAA tcc atg gaa aag aga aga tg | ||
| + | ***'''NO260''' = SPT3_fwd, GCT TTG ACC AAC TTT AAA GGT GGT AGA CTC AGT TCT AAA CCA ATT ATC ATG TAA, Tm = 64 | ||
| + | ***'''NO261''' = SPT7_fwd, TAC CAA TAT GGG CAG CAA TAG TTC ATT TAG CTT GAG CCT TCC TCG CCT TAA TCA ATA A, Tm = 67 | ||
| + | **Experimental primers for checking TAP tag | ||
| + | ***'''NO262''' = SGF73_fwd, ATT GAA GTA GGT ATT GGA AAT TCT GTG AAC CCC TAC AAT GGC AGA ATA AAT TAA, Tm = 64 | ||
| + | ***'''NO263''' = SGF29_fwd, GCC TTC GCC AAC GGC TTT GGC AAA CCT AGC AAG GAA ATA G, Tm = 68 | ||
| + | ***'''NO264''' = SGF11_fwd, ACT AGC AGC TCA TTT ACA GAG ATG TTT GAG TAG GGG TGC TAG ACG TTG A , Tm = 67 | ||
| + | ***'''NO265''' = UBP8_fwd, GGT TTT AAA GGA ACA GGC ATA TTT ATT ATT CTA CAC CAT TCG TCA AGT AAA TTG A, Tm = 63 | ||
| + | ***'''NO266''' = SUS1_fwd, GGG AAA CCG TTT TAA AGC AAA TAA GGG AAT TTC TTG AAG AGA TTG TAG ATA CAC AAT GA, Tm = 65 | ||
| + | ***'''NO277''' = RNT1_fwd, GAT CCC TCA CAA AAG AAT AAG AAA AGA AAA TTC TCA GAT ACA AGC TGA, Tm = 62 | ||
Latest revision as of 19:06, 28 July 2015
Contents
Introduction
To begin today���s experiment, you will ���pop��� some trp+ yeast from your transformation plates and then amplify the relevant portion of the released genomic DNA. The primers you will use are expected to give a ~554 base pair product only if the TAP-TRP tag is present. This is accomplished by using a forward primer that anneals to the very 5'-end of the TAP gene you've tried to add to your gene of interest and a reverse primer that anneals to a very 3'-end of the TRP gene that was part of the TAP tag and used for selection. There are two important caveats about today's experiment. First, if the TAP tag is fused elsewhere in the genome (giving rise to the trp+ phenotype) but the SAGA- or SAGA-controlled gene remains intact, then you may still see a product from this reaction. Thus we'll have to follow up in other ways to confirm that the gene of interest has, in fact, been modified. Second, you should be aware that a negative result from these reactions (i.e. no PCR product) can just as easily be explained as a failed PCR (bad primers, dead enzyme, wrong reaction conditions, etc) as a failed tagging transformation. So don't let a negative result today fool you and lead you to toss out perfectly correct samples. Only the positive result is meaningful in this experiment and we will not know the result until you run the agarose gel next time. So, we will remain optimistic and set up overnight cultures of both the candidates you are examining today. In this way you will have cells to examine next time, when you will check them for the TAP-tag by Western blot and when you will isolate total RNA from them for microarray analysis.Protocols
Part 1: Colony PCR
- Begin by counting the colonies that arose from the transformation experiment you performed last time. Choose two candidates (when possible) to pursue, circling the colonies on the back of the petri dish and clearly labeling them "A," and "B."If you do not have enough colonies, don't despair. You can get some additional ones from another group or from the teaching faculty. Note, however, that by doing so, you may be switching the gene you'll study for the rest of the module, and you will have to rethink the work you have planned so far. And if applicable, it will be worth noting the "negative" result in the research article you write, since it may indicate that despite the gene being deemed non-essential, cells may fare less well than wild-type cells when the gene is modified and so modified forms may be difficult to isolate.
- You will use the microwave to release the DNA from the yeast. On the tip of a sterile toothpick, pick-up 1/2 of the colony that is candidate "A" and swirl the cells in 20 μl of sterile water in an eppendorf tube. Be sure to label the tube so you know it belongs to your group and which candidate it contains. Repeat with the second candidate you've selected as well as with the trp+ version of the parental strain, NY420 and a positive control strain for this part of the experiment, either NY412 (same as NY411 but SPT3-TAP tagged) or NY413 (same as NY411 but SPT7-TAP tagged). The teaching faculty will have the control strains streaked out on petri dishes. You should mix 1/2 of a colony from these plate with 20 μl of sterile water as you did for your candidate colonies.
- Close the caps to the four tubes and microwave them in an eppendorf rack for 15 seconds.
- Move 2.5 μl of the microwaved mixes, yeast debris and all, into a 200 μl PCR tube. Again label your tubes well (write small!).
- Prepare your PCR cocktails. These are best performed with a reaction cocktail that contains the reagents common to all 4 reactions. In a new eppendorf tube (normal size this time), prepare a PCR cocktail enough for 5 reactions since you'll want a little extra volume to account for pipetting error. Each reaction will include:
- 10 μl 2.5X PCR mastermix
- 0.5 μl universal forward primer NO224, specific to the TAP portion of your tag
- 0.5 μl universal reverse primer NO225, specific to the TRP portion of your tag
- water to a final volume of 22.5 μl.
- Add 22.5 μl of PCR cocktail to each PCR tube with yeast and leave the tubes chilling until everyone is ready.
- Cycle the reactions as:
- 95° 4 minutes
- 95° 1 minute
- 52° 1 minute
- 72° 2 minutes
- repeat steps 2-4 35 times
- 72° 10 minutes
- 4° forever
Part 2: Overnight cultures
- Using sterile technique, aliquot 3 ml of YPD into four sterile test tubes.
- Label the caps with your team color, and either "parent" "+control" "A" or "B."
- Use sterile dowels to move the second 1/2 of each yeast colony from your transformation plates to the media. Swirl the dowel to remove the yeast from the stick and vortex the solution to fully resuspend them.
- Use a sterile dowel to innoculate the "parent" and "+control" cultures, using the NY420 and NY412 or NY413 plates that the teaching faculty have prepared.
- Leave your tubes with the teaching faculty who will place them on the roller drum in the 30�� incubator. The cultures will grow for 24 hours and then be placed in the 4�� fridge until next lab.
- When everyone is ready we will discuss the journal article that was assigned for today.
Part 3: Journal article discussion
As part of your assignment for today, you have read the relevant article by Wu et al, published in Mol Cell in 2004. There is also an associated review article [1] that was written to celebrate and highlight this important work on the structure of the SAGA complex. You and your partner will be randomly assigned a portion of the text to describe to the class but be prepared to engage in conversation about other parts of this paper even if you're not "assigned" to the section being discussed.
DONE!
For next time
- Write a paragraph to report on your transformation results. Eventually this homework assignment will be integrated into the results section for your research article that you'll hand in a few weeks from now. Report what you did and what you found here. Leave interpretation to the next FNT question.
- Write a sentence or two to interpret your transformation results. Eventually this information will be integrated into the discussion section of your research article that you'll hand in a few weeks from now.
- You are not required to hand in a revised Materials and Methods section, but consider adding the experiment you performed today to the assignment from last time.
- Next lab may run long!!! Plan ahead and come prepared with your strategy for testing phenotypes of your strains. What kinds of media are you interested in testing? If you'd like media besides YPD, YPGAL, -trp and -his, you must let us know one day in advance of lab so we can prepare them for you.
- Sign up for one of the two Journal Club presentations days here. We must split the class evenly between the two available days. First come first served. You do not have to select a paper to present yet (unless you want to!).
Reagents list
- YPD
- 10g yeast extract
- 20g peptone
- 20g glucose
- 20g agar (for liquid media, leave agar out!)
- add 1L water, autoclave 20min, stir to cool
- PCR reaction mix
- Primers
- Universal forward primer
- NO224 TCC ATG GAA AAG AGA AGA TGG AAA AAG AAT TTC ATA GCC GTC TC
- use with rev primer for checking TAP-TRP tag by PCR
- 44 mer
- Tm = 63.6
- Universal reverse primer
- NO225 TCA GGT TGA CTT CCC CGC GGA ATT CGC GTC TAC
- use with fwd primer for checking TAP-TRP tag by PCR
- expect length of TAP seq + TRP K. l seq = 554 bps (from 2147 to 2701 in pBS1479)
- Tm = 68
- Other positive control primers (not used today)
- NO258 for SPT3-TAP strain NY412, TTT AAA GGT GGT AGA CTC AGT TCT AAA CCA ATT ATC ATG tcc atg gaa aag aga aga tg
- NO259 for SPT7-TAP strain NY413, AAT AGT TCA TTT AGC TTG AGC CTT CCT CGC CTT AAT CAA tcc atg gaa aag aga aga tg
- NO260 = SPT3_fwd, GCT TTG ACC AAC TTT AAA GGT GGT AGA CTC AGT TCT AAA CCA ATT ATC ATG TAA, Tm = 64
- NO261 = SPT7_fwd, TAC CAA TAT GGG CAG CAA TAG TTC ATT TAG CTT GAG CCT TCC TCG CCT TAA TCA ATA A, Tm = 67
- Experimental primers for checking TAP tag
- NO262 = SGF73_fwd, ATT GAA GTA GGT ATT GGA AAT TCT GTG AAC CCC TAC AAT GGC AGA ATA AAT TAA, Tm = 64
- NO263 = SGF29_fwd, GCC TTC GCC AAC GGC TTT GGC AAA CCT AGC AAG GAA ATA G, Tm = 68
- NO264 = SGF11_fwd, ACT AGC AGC TCA TTT ACA GAG ATG TTT GAG TAG GGG TGC TAG ACG TTG A , Tm = 67
- NO265 = UBP8_fwd, GGT TTT AAA GGA ACA GGC ATA TTT ATT ATT CTA CAC CAT TCG TCA AGT AAA TTG A, Tm = 63
- NO266 = SUS1_fwd, GGG AAA CCG TTT TAA AGC AAA TAA GGG AAT TTC TTG AAG AGA TTG TAG ATA CAC AAT GA, Tm = 65
- NO277 = RNT1_fwd, GAT CCC TCA CAA AAG AAT AAG AAA AGA AAA TTC TCA GAT ACA AGC TGA, Tm = 62
- Universal forward primer