20.109(S10):Prepare RNA by IVT (Day3)
Contents
Introduction
So far you have prepared the DNA encoding both the 6-5 and the 8-12 aptamer fragments. However, it is the secondary structure of the RNA that actually allows the 8-12 sequence to bind to heme. In order to make RNA from DNA, you will perform an in vitro transcription (IVT) reaction.
What is needed to create RNA from DNA? In PCR, you used a DNA polymerase and dNTPs to make your DNA. In an IVT, you will use an RNA polymerase and NTPs instead. The polymerase is purified from T7… The buffer conditions are also somewhat different (check out the reagent lists at the end of each day), but both contain the important co-factor Mg2+. Finally, an IVT contains pyrophosphatase, because it has been empirically found to increase efficiency.
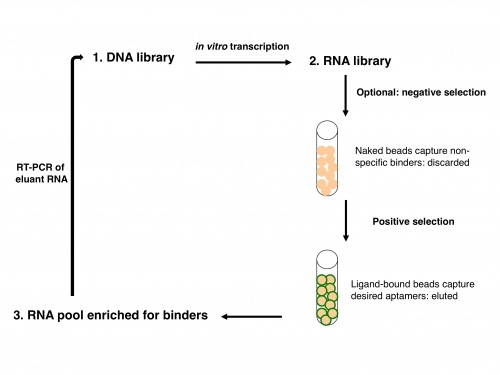
Now let's consider how an IVT fits into the overall scheme of SELEX. SELEX was simultaneously reported by two groups in 1990: Ellington and Szostak described RNA that bound to select aromatic small molecules, while Tuerk and Gold isolated RNA that bound to a DNA polymerase. As shown in the figure at right, due to the stability of DNA one usually starts with a DNA library rather than directly with an RNA library. The DNA is copied into RNA by transcription, and the resulting RNA pool is run over an affinity column that has the ligand of interest bound to it. Aptamers that do not bind are washed away by flowing buffer through the column. Afterward, RNA that does bind to the ligand is washed off, either by using free ligand as a competitor or sometimes by other changes in buffer conditions (e.g., change in salt concentration or pH). This RNA pool is copied into DNA by reverse transcription, then amplified by PCR, often using a one-step kit these days. After RT-PCR, the new library (presumably containing many fewer species of DNA) is again transcribed to RNA. Now the new library may be further refined by a second column selection.
An optional but often important step is to perform a negative selection. Incubating the aptamer pool with a column resin that lacks ligand but is otherwise identical to the affinity column can rid the pool of aptamers that bind non-specifically to the column material. If one is interested in finding an aptamer that binds to a particular ligand but not to several similar ligands, one might also perform negative selections using affinity columns with those undesired ligands bound.
After one or more column selections, one may use cloning to select individual RNA species for sequencing or for testing in a functional assay. In our case, the functional assay measures binding to heme by spectrophotometry. A typical starting library might have 10^13 to 10^15 sequences! In Szostak’s early binding experiments, molecules that bound to the aromatic dyes were enriched from less than 1% to greater than 50% of the RNA library population after four rounds of selection.
The IVT will run during the whole lab period… In the meantime, we will discuss a journal article, both to learn more about RNA aptamers and to become comfortable reading and discussing the primary scientific literature. In 2-3 weeks, you will each present an article on your own. Also have Atissa come?
Protocols
Because you are preparing RNA, you will have to take special precautions today and for the rest of the module. RNA is strikingly different from DNA in its stability. Consequently it is more difficult to work with RNA in the lab. It is not the techniques themselves that are difficult; indeed, many of the manipulations are nearly identical to those used for DNA. However, RNA is rapidly and easily degraded by RNases that exist everywhere. There are several rules for working with RNA. They will improve your chances of success. Please follow them all.
- Use warm water on a paper towel to wash lab equipment, like microfuges, before you begin your experiment. Then wipe them down with “RNase-away” solution.
- Wear gloves when you are touching anything that will touch your RNA.
- Change your gloves often.
- Before you begin your experiment clean your work area, removing all clutter. Wipe down the benchtop with warm water then “RNase-away,” and then lay down a fresh piece of benchpaper.
- Use RNA-dedicated solutions and if possible RNA-dedicated pipetmen.
- Start a new box of pipet tips and label their lid “RNase FREE.”
Part 1: In vitro transcription
The table below lists the amount of each reaction component needed for an 80 μL IVT. First, you should calculate how much total IVT Master Mix to make (2 or 4 rxns depending on whether you and your partner share one mix or not, plus 10% excess) and check your calculations with the teaching faculty if desired. Next, you can aliquot the appropriate amount of Master Mix into a number of eppendorf tubes, then add the relevant DNA to each labeled tube.
| Reagent | Amount for 1 reaction (μL) | Amount for N reactions + 10% |
|---|---|---|
| G7 buffer (2.5X stock) | 32 | |
| 1N KOH | 4.48 | |
| Pyrophosphatase | 4 | |
| NTPs | 22.4 | |
| T7 polymerase | 4 | |
| DNA | 13.1 | N/A |
Place your reaction tubes on the 37 °C heat block and write the time in your notebook. When everyone is ready, we will begin the journal article discussion. Also have Atissa come in and give her presentation talk?
- Put your reactions in the 37 °C heat block. After 3-4 hours, they will be frozen at – 20 °C until next time.
OR
- Put your reactions in the 37 °C heat block. When everyone is ready, we will have today’s lecture and quiz. After 3 hours, add 2 (per 20) μL DNase I to each reaction. After 30 min, they will be frozen at – 20 °C until next time.