Difference between revisions of "20.109(F07): M13.1"
| Line 2: | Line 2: | ||
==Introduction== | ==Introduction== | ||
| − | "Divide and conquer" may be an effective military strategy but its usefulness is not limited to that arena. The reductionist approach has been an important means of understanding complex biological processes. By tweezing apart networks and pathways, the components that contribute to the overall behavior of the system may be understood in great detail. As you've seen, however, | + | "Divide and conquer" may be an effective military strategy but its usefulness is not limited to that arena. The reductionist approach has been an important means of understanding complex biological processes. By tweezing apart networks and pathways, the components that contribute to the overall behavior of the system may be understood in great detail. As you've seen, however, there isn't always a straightforward way to reassemble the component level understanding into a predictive and quantitative model for the system. Remember that's what happened with the T7 model that you read about last time, and the author's response to the limited success of the models is what makes that T7 work so novel. Rather than continue to tweeze apart and better understand the natural example, they built a surrogate T7 that was a better template for experimental work, easier to manipulate and analyze, easier to characterize and understand. Now you will help design a surrogate M13. |
| + | |||
| + | [[Image:RefactorAlgortm.png|center| 600 px |T7.1 refactoring algorithm]] | ||
| + | |||
| + | To help with your refactoring efforts, a step-by-step example is provided here. It's inspired by the T7 algorithm described in the [http://www.nature.com/doifinder/10.1038/msb4100025 Chan et al paper.] The example provided here will almost certainly raise some but not all of the issues you will encounter as you try your own refactoring, so read it carefully and then use some or none of the example for your own work. | ||
| + | |||
| + | This example addresses the overlap in the genes encoding p9 and p8. In the natural version of M13K07, the promoter for gene 8 is not in a natural place and the stop codon for gene 9 overlaps the start codon for gene 8. This arrangement makes it nearly impossible to regulate expression of p9 and p8 independently---akin to the gas pedal of your car if it changed both the speed you travel and the volume of the radio. | ||
| + | |||
| + | ===Step 1: identify parts=== | ||
| + | As you've seen, many bacterial genes have definable "parts," i.e. genetic elements with particular purposes and without which the function of the DNA is impaired. The typical parts are: promoter, ribosome binding site, open reading frame, and terminator. In [http://www.ncbi.nlm.nih.gov/sites/entrez?Db=pubmed&Cmd=ShowDetailView&TermToSearch=6254849&ordinalpos=8&itool=EntrezSystem2.PEntrez.Pubmed.Pubmed_ResultsPanel.Pubmed_RVDocSum| the M13 sequencing paper] that was published in 1980, these parts were identified: | ||
| + | |||
| + | [[Image:Promoterseq Gene80.png|left|500 px|Identified promoter sequences]] | ||
| + | [[Image:Terminators gene80.png|right|500 px|Identified transcriptional terminators]] | ||
| + | [[Image:RBS gene80.png|left|300 px|Identified RBS sequences]] | ||
| + | |||
| + | Another useful resource is the [http://parts.mit.edu/registry/index.php/Main_Page Registry of Standard Biological Parts]. Many of the functional elements of the M13K07 genome have been entered into that database. A table describing them all is included in the "reagents list" for today's lab. | ||
| + | |||
| + | Armed with this information and the [http://www.neb.com/nebecomm/tech_reference/restriction_enzymes/sequences/m13ko7.txt| M13K07 sequence file], a working copy of the sequence can be copied into a new MSWord document and the relevant portions of the sequence annotated. The following color scheme was used and it would be very helpful if you could follow similarly when you refactor your portion of the genome | ||
| + | * RBS = blue box | ||
| + | * promoters = underlined with -35 and -10 in blue letters, bolding -10, -35 nucleotide | ||
| + | * transcriptional terminators = if relevant, boxing in grey | ||
| + | * as you box and color the features, you should note above region what’s what | ||
| + | |||
| + | [[Image:Refactor annotation.png|center| 300 px |Step 1: identify parts]] | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | Some of the identified genetic elements have been specified at the [http://parts.mit.edu/registry/index.php/Main_Page Registry of Standard Biological Parts]. A sample refactoring for M13, tweezing apart gene 8 and 9 is shown here as a model for you to follow. | ||
| + | |||
| + | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
[[Image:Gene8promoter?.png|thumb|promoter for gene 8?]] | [[Image:Gene8promoter?.png|thumb|promoter for gene 8?]] | ||
[[Image:PromotervsConsensus.png|thumb|Step 2: characterize parts]] | [[Image:PromotervsConsensus.png|thumb|Step 2: characterize parts]] | ||
| Line 17: | Line 44: | ||
| + | ===Part 1: M13.1=== | ||
| + | <center> | ||
| + | [[Image:Macintosh_HD-Users-nkuldell-Desktop-m13k07scaffold.png|600 px| M13K07 scaffold for re-engineering]] | ||
| + | </center> | ||
| + | Refactor the M13K07 genome from the unique HpaI site in gene II through the unique BamHI site in gene III, guided by the algorithm described in the Refactoring T7 paper [http://www.nature.com/msb/journal/v1/n1/suppinfo/msb4100025_S1.html Supplementary Figure 1]. You might start by annotating the M13K07 sequence across this region, giving hypothetical part names ("BBb_M13_109.07_00001") to all genetically encoded functions. Collect these parts in a table similar to the one in the introduction to today's lab and write a brief component descriptions for each. If there are particular aspects of the M13K07 sequence that cannot be edited, note these difficulties on your parts table, describing the design impass in sufficient detail that someone unfamiliar with our work might understand enough to help. | ||
| + | Proposed refinements can be noted on the individual parts as well as on our [[20.109(F07):Module 1:RefactorM13| M13 refactoring workpage]] | ||
| + | |||
| + | ===Part 2: Oral Presentation Instruction=== | ||
| + | Over the course of the term, everyone taking 20.109 will be making two presentations to the class. One will be a presentation of a primary journal article related to either [[20.109(F07): Module 1 oral presentations| genome]] or [[20.109(F07): Module 2 oral presentations| expression]] engineering. The other presentation will be of [[20.109(F07): Module 3 oral presentations| a research idea]]. Today our wonderful coach from the Writing program will come to lab to give a talk about giving talks. This presentation should be a welcomed chance to learn what's expected when you speak (in 20.109 and elsewhere) and to gather some great tips for giving polished and interesting seminars. You can follow up this instruction by re-reading the [[20.109(F07):Guidelines for oral presentations]]. | ||
| + | |||
| + | ===Part 3: Sequencing=== | ||
| + | The invention of automated sequencing machines has made sequence determination a fast and inexpensive endeavor. The method for sequencing DNA is not new but automation of the process is recent, developed in conjunction with the massive genome sequencing efforts of the 1990s. At the heart of sequencing reactions is chemistry worked out by Fred Sanger in the 1970’s which uses dideoxynucleotides. | ||
| + | <center> | ||
| + | [[Image:Be109dideoxynucleotide.jpg|thumb|left|500px|'''Normal bases versus chain-terminating bases''']] | ||
| + | </center> | ||
| + | <br style="clear:both" /> | ||
| + | |||
| + | These chain-terminating bases can be added to a growing chain of DNA but cannot be further extended. Performing four reactions, each with a different chain-terminating base, generates fragments of different lengths ending at G, A, T, or C. The fragments, once separated by size, reflect the DNA’s sequence. In the “old days” (all of 10 years ago!) radioactive material was incorporated into the elongating DNA fragments so they could be visualized on X-ray film (image on left). More recently fluorescent dyes, one color linked to each dideoxy-base, have been used instead. The four colored fragments can be passed through capillaries to a computer that can read the output and trace the color intensities detected (image on right). Your sample will be sequenced in this way on an ABI 3730 DNA Analyzer. | ||
| + | |||
| + | [[Image:Be109sequencinggel.jpg|thumb|left|120px|'''Sequencing gel''']] | ||
| + | [[Image:Be109tracedata.jpg|thumb|right|150px|'''Sequence trace data''']] | ||
| + | <br style="clear:both" /> | ||
| + | |||
| + | Sequencing will be one of the last steps to assess the success of your work on the gene for p3. From your experiment last time, you may feel confident that your oligonucleotide pair was successfully inserted into the M13K07 genome. What remains unclear, however is which direction the insert has gone in. Since the restriction sites are so close together, the orientations can't be distinguished by restriction analysis. To know if the construct you have in hand is in the intended or the reverse direction, it will be necessary to sequence the region. | ||
| + | |||
| + | Sequencing reactions require 200-500 ng of plasmid DNA, and 3.2 pmoles of sequencing primer in a final volume of 12 μl. The miniprep'd plasmid should have ~1 ug of nucleic acid/ul but that will be a mixture of RNA and DNA, so we will guess at the amount of plasmid DNA there and use 1 ul of your miniprep'd DNA/sequencing reaction. | ||
| + | |||
| + | Prepare an eppendorf tube with enough material for two sequencing reactions, namely | ||
| + | * 2 ul of your plasmid DNA candidate(s) | ||
| + | * 6.4 ul of a 1:100 dilution of the sequencing primer NO198 | ||
| + | * 15.6 ul sterile water | ||
| + | Pipet this solution up and down to mix and then transfer 12 ul to one of the tubes in the 8-tube strip we will turn in at the [http://web.mit.edu/biopolymers/www/| biopolymer's facility] in E17 for sequencing. The data will be available for you to examine one week from today. Keep your fingers crossed. | ||
| + | |||
| + | |||
| + | DONE! | ||
| + | |||
| + | ===For Next Time=== | ||
| + | # Exchange writing assignment with lab partner for comments/reactions. | ||
| + | # Prepare agarose gel figure. | ||
| + | |||
| + | ===Reagents list=== | ||
| + | |||
| + | * Sequencing primers | ||
| + | ** M13KO7seq_2101_fwd | ||
| + | ***NO198 5'- CAA GGC ACT GAC CCC GTT AAA ACT TAT TAC -3' | ||
| + | **M13KO7seq_2151_fwd | ||
| + | ***NO199 5'- AAA AGC CAT GTA TGA CGC TTA CTG GAA CGG -3' | ||
| + | * Genetic parts for M13K07 in the [http://parts.mit.edu/registry/index.php/Main_Page| Registry of Standard Biological Parts] | ||
<center> | <center> | ||
{| border="1" | {| border="1" | ||
| Line 91: | Line 166: | ||
</center> | </center> | ||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
Revision as of 16:01, 26 August 2007
Contents
Introduction
"Divide and conquer" may be an effective military strategy but its usefulness is not limited to that arena. The reductionist approach has been an important means of understanding complex biological processes. By tweezing apart networks and pathways, the components that contribute to the overall behavior of the system may be understood in great detail. As you've seen, however, there isn't always a straightforward way to reassemble the component level understanding into a predictive and quantitative model for the system. Remember that's what happened with the T7 model that you read about last time, and the author's response to the limited success of the models is what makes that T7 work so novel. Rather than continue to tweeze apart and better understand the natural example, they built a surrogate T7 that was a better template for experimental work, easier to manipulate and analyze, easier to characterize and understand. Now you will help design a surrogate M13.
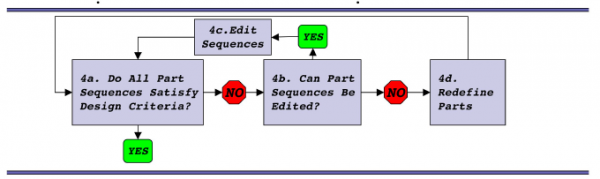
To help with your refactoring efforts, a step-by-step example is provided here. It's inspired by the T7 algorithm described in the Chan et al paper. The example provided here will almost certainly raise some but not all of the issues you will encounter as you try your own refactoring, so read it carefully and then use some or none of the example for your own work.
This example addresses the overlap in the genes encoding p9 and p8. In the natural version of M13K07, the promoter for gene 8 is not in a natural place and the stop codon for gene 9 overlaps the start codon for gene 8. This arrangement makes it nearly impossible to regulate expression of p9 and p8 independently---akin to the gas pedal of your car if it changed both the speed you travel and the volume of the radio.
Step 1: identify parts
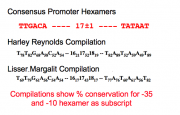
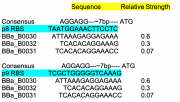
As you've seen, many bacterial genes have definable "parts," i.e. genetic elements with particular purposes and without which the function of the DNA is impaired. The typical parts are: promoter, ribosome binding site, open reading frame, and terminator. In the M13 sequencing paper that was published in 1980, these parts were identified:
Another useful resource is the Registry of Standard Biological Parts. Many of the functional elements of the M13K07 genome have been entered into that database. A table describing them all is included in the "reagents list" for today's lab.
Armed with this information and the M13K07 sequence file, a working copy of the sequence can be copied into a new MSWord document and the relevant portions of the sequence annotated. The following color scheme was used and it would be very helpful if you could follow similarly when you refactor your portion of the genome
- RBS = blue box
- promoters = underlined with -35 and -10 in blue letters, bolding -10, -35 nucleotide
- transcriptional terminators = if relevant, boxing in grey
- as you box and color the features, you should note above region what’s what
Some of the identified genetic elements have been specified at the Registry of Standard Biological Parts. A sample refactoring for M13, tweezing apart gene 8 and 9 is shown here as a model for you to follow.
Media:TheBacterialPromoter.pdf downloaded refresher on bacterial promoters
Part 1: M13.1
Refactor the M13K07 genome from the unique HpaI site in gene II through the unique BamHI site in gene III, guided by the algorithm described in the Refactoring T7 paper Supplementary Figure 1. You might start by annotating the M13K07 sequence across this region, giving hypothetical part names ("BBb_M13_109.07_00001") to all genetically encoded functions. Collect these parts in a table similar to the one in the introduction to today's lab and write a brief component descriptions for each. If there are particular aspects of the M13K07 sequence that cannot be edited, note these difficulties on your parts table, describing the design impass in sufficient detail that someone unfamiliar with our work might understand enough to help.
Proposed refinements can be noted on the individual parts as well as on our M13 refactoring workpage
Part 2: Oral Presentation Instruction
Over the course of the term, everyone taking 20.109 will be making two presentations to the class. One will be a presentation of a primary journal article related to either genome or expression engineering. The other presentation will be of a research idea. Today our wonderful coach from the Writing program will come to lab to give a talk about giving talks. This presentation should be a welcomed chance to learn what's expected when you speak (in 20.109 and elsewhere) and to gather some great tips for giving polished and interesting seminars. You can follow up this instruction by re-reading the 20.109(F07):Guidelines for oral presentations.
Part 3: Sequencing
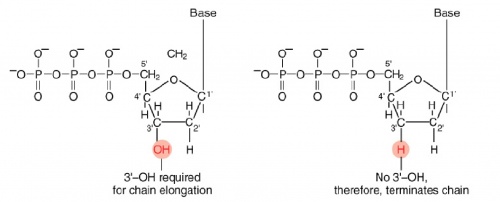
The invention of automated sequencing machines has made sequence determination a fast and inexpensive endeavor. The method for sequencing DNA is not new but automation of the process is recent, developed in conjunction with the massive genome sequencing efforts of the 1990s. At the heart of sequencing reactions is chemistry worked out by Fred Sanger in the 1970’s which uses dideoxynucleotides.
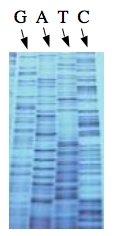
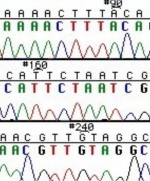
These chain-terminating bases can be added to a growing chain of DNA but cannot be further extended. Performing four reactions, each with a different chain-terminating base, generates fragments of different lengths ending at G, A, T, or C. The fragments, once separated by size, reflect the DNA’s sequence. In the “old days” (all of 10 years ago!) radioactive material was incorporated into the elongating DNA fragments so they could be visualized on X-ray film (image on left). More recently fluorescent dyes, one color linked to each dideoxy-base, have been used instead. The four colored fragments can be passed through capillaries to a computer that can read the output and trace the color intensities detected (image on right). Your sample will be sequenced in this way on an ABI 3730 DNA Analyzer.
Sequencing will be one of the last steps to assess the success of your work on the gene for p3. From your experiment last time, you may feel confident that your oligonucleotide pair was successfully inserted into the M13K07 genome. What remains unclear, however is which direction the insert has gone in. Since the restriction sites are so close together, the orientations can't be distinguished by restriction analysis. To know if the construct you have in hand is in the intended or the reverse direction, it will be necessary to sequence the region.
Sequencing reactions require 200-500 ng of plasmid DNA, and 3.2 pmoles of sequencing primer in a final volume of 12 μl. The miniprep'd plasmid should have ~1 ug of nucleic acid/ul but that will be a mixture of RNA and DNA, so we will guess at the amount of plasmid DNA there and use 1 ul of your miniprep'd DNA/sequencing reaction.
Prepare an eppendorf tube with enough material for two sequencing reactions, namely
- 2 ul of your plasmid DNA candidate(s)
- 6.4 ul of a 1:100 dilution of the sequencing primer NO198
- 15.6 ul sterile water
Pipet this solution up and down to mix and then transfer 12 ul to one of the tubes in the 8-tube strip we will turn in at the biopolymer's facility in E17 for sequencing. The data will be available for you to examine one week from today. Keep your fingers crossed.
DONE!
For Next Time
- Exchange writing assignment with lab partner for comments/reactions.
- Prepare agarose gel figure.
Reagents list
- Sequencing primers
- M13KO7seq_2101_fwd
- NO198 5'- CAA GGC ACT GAC CCC GTT AAA ACT TAT TAC -3'
- M13KO7seq_2151_fwd
- NO199 5'- AAA AGC CAT GTA TGA CGC TTA CTG GAA CGG -3'
- M13KO7seq_2101_fwd
- Genetic parts for M13K07 in the Registry of Standard Biological Parts
| genetic element | promoter | RBS | coding |
|---|---|---|---|
| arbitrary starting point | |||
| gII | BBa_M13102 | BBa_M13502 | BBa_M13002 |
| gX | BBa_M13110 | BBa_M13510 | BBa_M13010 |
| gV | BBa_M13105 | BBa_M13505 | BBa_M13005 |
| gVII | BBa_M13507 | BBa_M13007 | |
| gIX | BBa_M13509 | BBa_M13009 | |
| gVIII | BBa_M13108 | BBa_M13508 | BBa_M13008 |
| Transcriptional terminator | |||
| gIII | BBa_M13103 | BBa_M13503 | BBa_M13003 |
| gVI | BBa_M13106 | BBa_M13506 | BBa_M13006 |
| gI | BBa_M13101 | BBa_M13501 | BBa_M13001 |
| gXI | BBa_M13511 | BBa_M13011 | |
| gIV | BBa_M13104 | BBa_M13504 | BBa_M13004 |
| M13K07 ori/KanR/p15a ori | |||