Assignment 10, Part 1: Measuring the osmotic shock response of yeast
Add a power saving module to your pinch valve circuit
You may have noticed in Assignment 8 that the pinch valves get quite hot if they are engaged for a long time. This is a problem because when the tubing gets hot, air bubbles can form in the medium. Air bubbles will disrupt your flow and rip cells off of the coverslip. We want to avoid air bubbles at all costs. The yeast cells respond to osmotic shock at very low frequencies (on long time scales), so we can't avoid leaving the valve engaged for long times. We'll mitigate this problem by inserting a power saving module to our valve-control circuit.
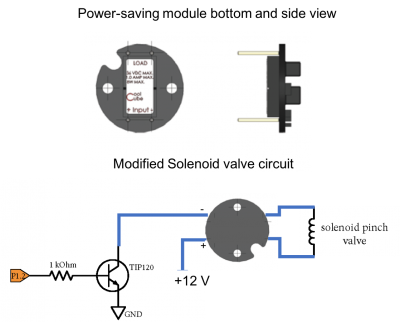
To engage the solenoid valve, we need power delivered from a +12V power supply. Once the solenoid is engaged, however, it requires much less power to remain engaged. The power module shown in the figure below is designed to supply a +12V signal for 100ms, then it drops the power delivered to the solenoid by about 1/3. In this state, the valve will still stay open, but it will produce much less heat (and fewer bubbles!).
- Modify your pinch valve circuit to use the power module as shown in the figure. Be careful to connect the +12V to the appropriate pin!
- Test that your computer controls still work for the pinch valve and LED illuminators.
Assemble a microfluidic device
Follow the instructions in Assignment 8 to assemble a PDMS/tape device. Note that you should still have your slab of cured PDMS left over from Assignment 8, so you don't have to repeat that step.
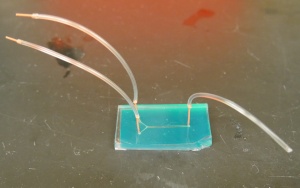
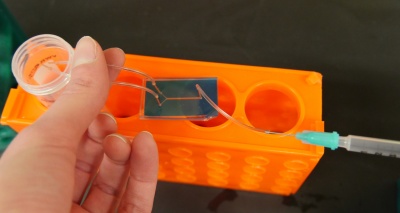
We'll cut some new tubing for this experiment, rather than reusing the tubing from Assignment 8. We want to be able to wash the flow channel with ethanol, water and ConA solutions, so we'll connect some short sections of tubing to the device before hooking it up to the media reservoirs (see figure on the right).
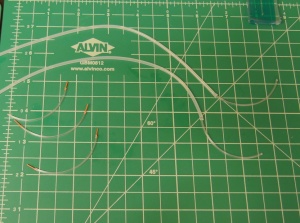
- Prepare two sections of device inlet tubing:
- Cut 2x sections of 2.5" lengths of tygon tubing.
- Cut 4x 1 cm lengths of PEEK tubing, and sleeve one of them into each end of short tygon tubing sections.
- Connect these to the inlet holes of your PDMS device.
- Prepare one section of device outlet tubing:
- Cut one 2.5" length of tygon tubing.
- Cut one 1 cm length of PEEK tubing, and sleeve it into the short section of tygon tubing.
- Connect the PEEK tubing to the outlet hole of your PDMS device.
- Prepare 2x sections of silicone/tygon tubing for media reservoirs:
- Cut 2x sections of 2.5" lengths of tygon tubing.
- Cut 2x 12" lengths of the flexible silicone tubing.
- Sleeve each section of silicone tubing onto one section of 2.5" tygon tubing.
- Prepare one section of waste reservoir tubing:
- Cut one 1 cm section of orange PEEK tubing and sleeve it into a 15" length of tygon tubing.
Incubate device with ConA
We need to immobilize yeast cells to the coverslip of the flow device in order to image them over long times. A standard method of immobilizing yeast cells for microscopy is to coat the coverslip with concanavalin A (ConA) - a lectin that binds to cell surface saccharides.
- Thaw 1 aliquot of ConA (250 uL of 1 mg/ml in PBS), preferably slowly, on ice.
- Wash your flow channel with 70% ethanol
- Load a 1 ml syringe with about 500 uL of 70% ethanol.
- Push the ethanol backwards through the device, from the outlet to the inlet, holding the inlets over a wast container.
- Next, wash with ~500 uL DI water.
- Load the ConA solution into a 1 ml syringe. Without introducing any bubbles, push the entire aliquot of ConA through the device. Leave the syringe connected an let incubate for at least 20 minutes (or until ready to load the yeast cells).
Set up media reservoirs and prime inlet tubing
It's common in fluidics experiments to "prime the lines" which means to fill up the tubing with liquid before connecting your microfluidic device. This step can help to prevent bubbles from getting lodged in your device. We want to avoid bubbles at all costs!
- Insert the 15 ml high and low salt media reservoirs into the 50 ml conical tube holders.
- Feed the silicone end of the inlet tubing into the high-salt reservoir. Make sure the end of the tube is at the bottom of the reservoir. Consider using lab tape to prevent it from getting dislodged.
- Connect a 1 ml syringe to the tygon end of the tubing using a 23G blunt-tipped needle, and pull the fluid from the reservoir all the way to the end of the tubing. Leave the syringe connected.
- Engage the solenoid valve (i.e. foo.OpenHighSalt), then feed the silicone tubing into the now-open slot of the pinch valve. Check that the tubing is completely seated inside the valve.
- Close the pinch valve (foo.OpenLowSalt) to prevent flow from the high-salt reservoir, then disconnect the needle and syringe.
- Repeat steps 2-3 with the low-salt reservoir. Feed the silicone tubing into the low salt slot of the pinch valve. Leave the syringe connected until ready to connect the PDMS device.
- If you're still waiting for the ConA to finish incubating, this may be a good time to check your microscope alignment using a reference slide.
Load yeast cells into the flow channel
The final setup steps include connecting the fluidic device to the reservoirs, then loading the yeas cells into the device. Care should be taken to avoid introducing bubbles, or applying too much pressure once the yeast cells are immobilized.
Connect device to fluid reservoirs
- Attach the high salt reservoir tubing to one inlet tube of the PDMS device.
- Engage the pinch valve to open the high salt valve and close the low salt valve.
- Disconnect the syringe from the low salt tubing and connect it to the remaining inlet tubing of the device.
- Connect the syringe to the outlet of the device, and (gently!) pull some fluid from the low salt reservoir through the device.
- Engage the valve to open the high salt reservoir and pull some of the high salt medium through the device.
- Toggle the valves several times and try to remove any visible bubbles by (gently!) pulling on the syringe.
- Leave the valve in the disengaged (low salt) state, and keep the syringe connected while you prepare the cells.
Prepare yeast cells
- Remove 2x 0.75 mL of the cell culture into two microcentrifuge tubes.
- The cells are grown in synthetic-complete medium (SC) which has a lower autofluorescence than other types of medium.
- The ideal yeast concentration for log-growth is for it to have an OD600 between 0.4 and 0.6.
- Add 0.75 ml of YPD medium to each culture tube. (YPD medium helps the cells form a better pellet when centrifuged.)
- Put both tubes in the microcentrifuge (so that they're balanced!) and spin at 800g for 2 minutes.
- Pipette off the supernatant and resuspend one of the pellets in 125 uL of SC medium. Combine the suspended culture with the second pellet, and resuspend it in the same 125uL solution.
Load the yeast cells into fluidic device
- Double check that the tubing and device are bubble free, and that the pinch valve is in the low salt state.
- Attach a 23G blunt-tipped needle to a syringe, and withdraw the yeast cells into it. Ensure that there is no air in the tip of the needle by pushing out a drop or two of the culture.
- Disconnect the syringe at the outlet of the PDMS device, and, without introducing any air bubbles, connect the syringe containing the yeast cells.
- Slowly inject the yeast cells into the PDMS device. Avoid introducing air into the tubing by pushing the plunger only 2/3 of the way.
- Leave the syringe connected, and let the cells settle and adhere to the coverslip for 10-15 minutes. (You can jump ahead to Find the focus and set the imaging parameters if you want to while you wait.)
- When ready, disconnect the syringe and connect the tygon outlet tubing in its place. Put the other end of the outlet tubing in the waste reservoir.
- You should see the meniscus of the media flow through the outlet tubing, ultimately forming droplets into the waste reservoir. If you can't see any flow, try raising the inlet reservoirs. As a last resort, connect a syringe to the end of the oultet tubing and SLOWLY withdraw (pulling too quickly may dislodge all your cells!).
- Toggle the pinch valve several times to ensure that both media reservoirs are flowing as expected, then leave the cells to equilibrate at low salt.
Record movies
Once you're sure the flow is behaving as expected, let the yeast flow under low salt conditions while you align your microscope and prepare to record data.
Find the focus and set the imaging parameters
- Check the alignment of both blue and green illumination for your microscope using a reference slide.
- Clip the flow channel onto the stage (use a red acrylic ring as support), and use the room lights (or high gain setting) to bring the yeast cells into focus.
- Cover your flow channel with a box to block out background light (careful that the box doesn't pull on the tubing).
- Turn the blue light on, and using a high gain setting, focus on the cells.
- Turn on the green excitation, and again using high gain, adjust the focus. The cell nuclei are harder to focus on, so make sure to always use the green illumination as your final focusing stage. If you don't see a good nucleus image, there's no hope for collecting good data.
- Lock the stage in all three directions using the locking screws. This is extremely important for long experiment times to prevent the stage from drifting out of focus.
- With the green illumination on, use the SetGreenImageParameters command to set the exposure time to 5s (5000000 us) and adjust the gain to get the best image.
- Turn on the blue illuminator and set the exposure and gain. In both cases, exposure times longer than 5 or 6 start to become impractical, so we can only increase gain at that point.
Start recording data!
Each group will be assigned three conditions to measure. The options are:
- T = 2 min. Record 5 periods, and acquire 4 images per period, equilibrate for just under 8 minutes (e.g. 478s).
- T = 4 min. Record 4 periods, and acquire 8 images per period, equilibrate for just under 8 minutes.
- T = 8 min. Record 3 periods, and acquire 8 images per period, equilibrate for just under 8 minutes.
- T = 16 min. Record 2 periods, and acquire 8 images per period, equilibrate for just under 16 minutes.
- T = 32 min. Bonus points if you complete this one. Record 2 periods, wait for 5-10 minutes at low salt before starting this experiment.
- Step response. Record for 2 minutes at low salt and 25 minutes at high salt. Acquire an image every 1 min.
Tips:
- Start with the smallest period and work your way up.
- These experiments are tricky and will only work if you are careful and vigilant. It's tempting to walk away while your data is recording, but check in frequently to make sure that everything is still behaving as expected. There's no use taking a 30 minute movie if your cells drifted out of focus in the first frame!
- Re-check your focus and camera settings after each measurement. Use high gain settings when focusing, long exposure times when taking data. You may also want to move your FOV to image fresh cells if the current ones seem dim.
- Don't forget to save your data to a variable in the workspace and SAVE YOUR WORKSPACE FREQUENTLY!
- Lock the z-axis of your stage when taking data to prevent drift in the focus of your microscope.
- The following are some useful commands:
% Initialize the software foo = OsmoticShocker; foo.Initialize; % Turn on and off the illumination (only one will be on at a time) foo.BlueOn foo.BlueOff foo.GreenOn foo.GreenOff % Set imaging parameters foo.SetBlueImageParameters(2,5000000) % for Gain = 2, exposure = 5s foo.SetGreenImageParameters(15, 5000000) % Set parameters for oscillation experiment foo.SetOscillationParameters % will prompt you for inputs foo.StartOscillationExperiment % will start oscillating the valves and will start recording images after the specified equilibration time % Set parameters for step response experiment foo.SetStepResponseParameters % will prompt you for inputs foo.StartStepExperiment
Clean up
When you're sure you have collected and saved all your data, follow these steps to clean up.
- Wearing gloves, remove the two inlet tubings from their reservoirs.
- Remove the silicone tubing from the pinch valve, and let the fluid drain through the device into the waste container.
- Disconnect the tubing from the PDMS device, dispose of the tubing in a biowaste container, and dispose of the device in the biological sharps bin.
- Aspirate the remaining liquid from the reservoirs and waste container, and dispose of the empty tubes in the biological waste bin.
- Discard any opened syringes and needles in the biological sharps waste.
Analyze data
Use your code from assignment 9 to analyze your movies.
Tips
- Run your analysis on the first frame of each movie to check that it is behaving as expected. Then proceed to analyze the whole movie.
- Analyzing the movies can be slow. Save your resulting Hog1-response to a .mat file to avoid having to run the movie analysis multiple times. For example:
save Hog1Response2min hog1response
will save only the variable hog1response to the file Hog1Response2min.mat.