Difference between revisions of "20.109(F16):Induce CRISPRi system (Day7)"
Noreen Lyell (Talk | contribs) (→Navigation links) |
Noreen Lyell (Talk | contribs) |
||
| (45 intermediate revisions by 3 users not shown) | |||
| Line 4: | Line 4: | ||
==Introduction== | ==Introduction== | ||
| − | + | The CRISPRi system involves three genetic elements: the target gene within the host genome, the pdCas9 plasmid, and the pgRNA_target plasmid. Though the target genome is native to the host cell, the plasmids must be transformed into the cell and maintained using antibiotic selection. Throughout this module, we have learned about and worked with the CRISPRi plasmids as individual units, but now we will consider the system as a whole in the context of gene regulation. | |
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | In the previous laboratory session, you co-transformed pdCas9 and your pgRNA_target plasmids into ''E. coli'' MG1655. The colonies present on your LB plates containing chloramphenicol and ampicillin should carry both plasmids in that they were able to survive selection by both antibiotics. The promoter (pJ23119) driving expression of your gRNA sequence in the gRNA_target plasmid is constitutively active. This means that RNAP is not prohibiting from binding and transcription of the gRNA target sequence is therefore not inhibited. Therefore, your gRNA_target is always present in the MG1655 cells and binding to the target sequence within the genome. | |
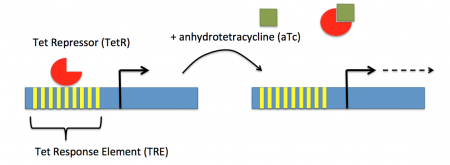
| − | + | [[Image:Fa16 M2D7 tet repression schematic.png|thumb|right|450px|'''Schematic of aTc induction of tet promoter.''']]In contrast, expression of the gene encoding Cas9 within pdCas9 is regulated by an inducible promoter (p<sub>L</sub>tetO-1). An inducible promoter is 'off' unless the appropriate molecule is present to relieve repression. In the case of our system, expression of the gene encoding Cas9 is inhibited due to the use of a tet-based promoter construct. Tet is shorthand for tetracycline, which is an antibiotic that inhibits protein synthesis through preventing the association between charged aminoacyl-tRNA molecules and the A site of ribosomes. Bacterial cells that carry the tet resistance cassette are able to survive exposure to tetracycline by expressing genes that encode an efflux pump that 'flushes' the antibiotic from the bacterial cell. To conserve energy, the tet system is only expressed in the presence of tetracyline. In the absence of tetracycline, a transcription repressor protein (TetR) is bound to the promoter upstream of the tet resistance cassette genes. When tetracycline is present, the molecule binds to TetR causing a confirmational resulting in TetR 'falling off' of the promoter. In the CRISPRi system, the tet-based promoter construct upstream of the gene that encodes Cas9 is 'off' unless anhydrotetracyline (aTc), an analog of tetracyline, is added to the culture media. | |
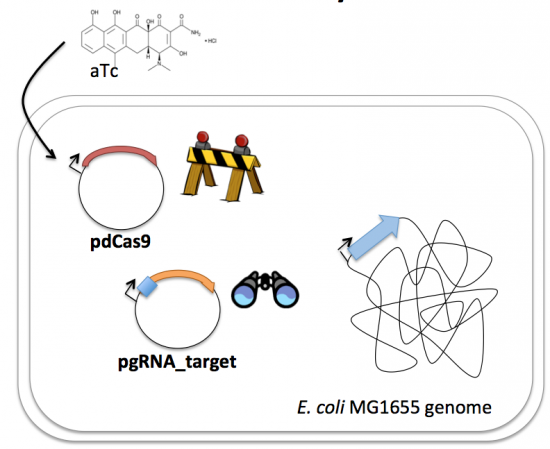
| − | + | Taken together, the gRNA-target molecule is constitutively transcribed and, as stated above, always present / binding to the target. The dCas9 protein is only present when aTc is added. Thus, gene expression is only altered when aTc is present. As represented by the schematic below, the gRNA_target 'seeks out' the target within the host genome and recruits dCas9 to the site. When associated with the target / gRNA_target complex, dCas9 binds to the site and acts as a 'roadblock' by prohibiting RNAP access to the sequence. Because the gene of interest is not able to be transcribed, the protein encoded by that gene is not synthesized. In our experiments, we hypothesize that the absence of specific proteins, or enzymes, in the fermentation pathway will increase the yield of either ethanol or lactate. | |
| + | |||
| + | [[Image:Fa16 M2D7 CRISPRi overview v4.png|thumb|center|550px|'''Schematic of CRISPRi system induction.''' Following addition of aTc, the dCas9 protein is produced and recruited by the gRNA target sequence to the target gene. Once recruiting, the dCas9 protein associates with the gRNA/genome complex and impedes transcription by RNAP.]] | ||
==Protocols== | ==Protocols== | ||
===Part 1: Communication Lab workshop=== | ===Part 1: Communication Lab workshop=== | ||
| − | + | Our communication instructor Dr. Sean Clarke will join us today for a workshop on organizing and writing your M2 research article. | |
| − | ===Part 2: Examine | + | ===Part 2: Examine pgRNA sequencing results=== |
| − | Your goal today is to analyze the sequencing data for you two potential mutant | + | Your goal today is to analyze the sequencing data for you two potential mutant pgRNA clones - two independent colonies from your amplification reaction - and then decide which colony to proceed with for the CRISPRi manipulation of the ''E. coli'' MG1655 fermentation pathway. |
| − | + | '''Retrieve sequence results from Genewiz''' | |
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | #Use the pgRNA_target Benchling file that you generated on [[20.109(F16):Confirm_gRNA_sequence_(Day5)#Part_3:_Prepare_pgRNA_clones_for_sequencing_analysis| M2D5]] to confirm that the gRNA sequence you designed was indeed inserted into the vector. | |
| + | #Your sequencing data is available from [http://genewiz.com Genewiz]. It was uploaded to the [http://engineerbiology.org/wiki/Talk:20.109(F16):Module_2 Module 2 main page Discussion] tab. | ||
| − | + | You should align your sequencing data with a known sequence, in this case the gRNA target sequence you selected, to identify any unintended base changes that may have occurred. There are several web-based programs for aligning sequences and still more programs that can be purchased. The steps for using Benchling are below. Please feel free to use any program with which you are familiar. | |
| − | + | ||
| − | + | '''Align with A Plasmid Editor'''<br> | |
| − | + | You can use any DNA manipulation software you choose to complete the alignment confirmation, but the instructions provided are for APE (A Plasmid Editor, created by M. Wayne Davis at the University of Utah). The software can be downloaded free-of-charge from [http://bioweb.biology.utah.edu/jorgensen/wayned/ape/ this site] onto your personal computer or you can use the 20.109 laboratory computers. Please note that if you use a different program the teaching faculty may not be able to assist you. | |
| − | + | ||
| − | #Paste the sequence text from your sequencing run into the | + | #Make an APE DNA file that contains the gRNA oligo you used on [[20.109(F16):Generate gRNA plasmid (Day3)| M2D3]]. This sequence should contain the target sequence you selected and the dCas9 tag sequence. |
| + | #*Open APE then copy and paste the sequence into a new workspace. | ||
| + | #Open a new window and create an APE DNA file that contains the results from your sequencing reaction. | ||
| + | #*Go to File and select 'New' to open a new window. | ||
| + | #Paste the sequence text from your sequencing run into the new window. If there were ambiguous areas of your sequencing results, these will be listed as "N" rather than "A" "T" "G" or "C" and it's fine to include Ns in the query. | ||
#*The start and end of your sequencing may have several Ns. In this case it is best to omit these Ns by pasting only the 'good' sequence that is flanked by the ambiguous sequence. | #*The start and end of your sequencing may have several Ns. In this case it is best to omit these Ns by pasting only the 'good' sequence that is flanked by the ambiguous sequence. | ||
| − | # | + | #Go to Tools and select 'Align Two Sequences...' |
| − | #Click | + | #*In one drop-down window choose the gRNA oligo file and in the second drop-down window choose the new file that contains the Genewiz sequence you copied and pasted in step #3. |
| − | #Carefully examine the sequence to see if your | + | #*Be sure to consider whether you want to compare the reverse-complement of the Genewiz sequence and, if appropriate, check the box to the right of the drop-down window. '''If you are unsure if this box should be checked, ask the teaching faculty.''' |
| − | #You should save a screenshot of each alignment and attach them to your notebook. | + | #Click 'OK' and a new window should open with the sequences aligned. Matches will be shown by vertical lines between the aligned sequences. You should see a long stream of matches. If your insertion is present, then in this stream of matches the inserted basepairs should be highlighted in red. |
| − | #Follow the above steps to examine all of your sequencing results. '''Remember: you used a forward and a reverse primer to interrogate both | + | #Carefully examine the sequence to see if your gRNA insertion was incorporated. |
| + | #You should save a screenshot of each alignment and attach them to your Benchling notebook. | ||
| + | #Follow the above steps to examine all of your sequencing results. '''Remember: you used a forward and a reverse primer to interrogate both potential gRNA_target plasmids.''' | ||
| + | |||
| + | If both clones for your gRNA_target have the correct sequence, choose either co-transformant to use for the aTc induction step. If only one is correct, then this is the co-transformant you will use next time. If neither of your plasmids carry the appropriate insertion, talk to the teaching faculty. | ||
| + | <!-- | ||
| + | '''Align with Benchling''' | ||
| + | #Make a Benchling DNA file that contains the gRNA oligo you used on [[20.109(F16):Generate gRNA plasmid (Day3)| M2D3]]. This sequence should contain the target sequence you selected and the dCas9 tag sequence. | ||
| + | #*Select the create button (plus sign) at the top of the Project window. | ||
| + | #*Select DNA. | ||
| + | #*Enter the gRNA oligo sequence. | ||
| + | #*Type 'gRNA oligo' into the Name text box. | ||
| + | #*Click "UPDATE INFORMATION" button. | ||
| + | #Download the Trace File for your forward and reverse sequences to your computer. | ||
| + | #*Create a folder for the files. | ||
| + | #Import these files into Benchling. | ||
| + | #*You need to be in the Inventory tab. | ||
| + | #*Select the create button (plus sign) at the top of the Project window. | ||
| + | #*Select Import DNA. | ||
| + | #*In the Convert Files tab, click the "OR CHOOSE A FILE" button. | ||
| + | #*Select the Trace Files from the folder into which you downloaded them above. | ||
| + | #*Before you continue, be sure the forward and reverse files are in the Inventory tab of your Project window. | ||
| + | #Find the consensus sequence for the forward and reverse sequences. | ||
| + | #*Select the expand button (right arrow) at the top of the Project window. | ||
| + | #*Check the boxes to the left of the forward and reverse sequence files. | ||
| + | #*Click the "More" button at the top right, then hover over Analyze, and select Create Alignment. | ||
| + | #*In the Alignment confirm that the forward and reverse sequence files are listed under the Sequences header. | ||
| + | #*Be sure MAFFT algorithm is selected under the Algorithm header. | ||
| + | #*Click "CREATE ALIGNMENT" button. | ||
| + | #*Click the Collapse button at the top right to view your alignment. | ||
| + | #Check the consensus sequence for mutations. | ||
| + | #*Red bars indicate the presence of a discrepancy between the forward and reverse sequences. If your alignment contains a discrepancy, check the traces at that location and attempt to resolve the issue by examining the quality of each sequence. | ||
| + | #*If you are unsure, please consult the teaching faculty. | ||
| + | #Save the consensus file. | ||
| + | #*Select the Sequence info button (i) at the right of the window that contains the consensus sequence. | ||
| + | #*Type 'pgRNA_target consensus' into the Name text box. | ||
| + | #*Click "UPDATE INFORMATION" button. | ||
| + | #Align gRNA oligo with consensus sequence. | ||
| + | #*Select the expand button (right arrow) at the top of the Project window. | ||
| + | #*Check the box to the left of the consensus file. | ||
| + | #*Check the box to the left of the gRNA oligo DNA file created in Step #1. | ||
| + | #*Click the "More" button at the top right, then hover over Analyze, and select Create Alignment. | ||
| + | #*In the Alignment confirm that the consensus and gRNA sequence files are listed under the Sequences header. | ||
| + | #*Be sure Clustal Omega algorithm is selected under the Algorithm header. | ||
| + | #*Click "CREATE ALIGNMENT" button. | ||
| + | #*Click the Collapse button at the top right to view your alignment. | ||
| + | #Confirm that the gRNA oligo sequence is present in your consensus sequence. | ||
| + | #Save the gRNA confirmation file. | ||
| + | #*Select the Sequence info button (i) at the right of the window that contains the consensus sequence. | ||
| + | #*Type 'pgRNA_target confirmation' into the Name text box. | ||
| + | #*Click "UPDATE INFORMATION" button. | ||
| − | If both | + | If both clones for your pgRNA have the correct sequence, choose one to use for the co-transformation step. If only one is correct, then this is the one you will use next time. If neither of your plasmids carry the appropriate gRNA target sequence, talk to the teaching faculty. |
| + | --> | ||
===Part 3: Prepare media for mixed-acid fermentation inoculations=== | ===Part 3: Prepare media for mixed-acid fermentation inoculations=== | ||
| + | To test the effect of your gRNA on altering the production of either ethanol or lactate, you will incubate the co-transformed ''E. coli'' cells both with and without oxygen. Remember from lecture that cells grown anaerobically will produce more fermentation products; however, our goal is to further enhance the production of these products by manipulating the fermentation pathway. | ||
| + | #Acquire 4 glass culture tubes and 4 15 mL conical tubes from the front laboratory bench and label as shown below: | ||
| + | #*MG1655 +O<sub>2</sub> -aTc (glass tube) | ||
| + | #*MG1655 -O<sub>2</sub> -aTc (15 mL conical tube) | ||
| + | #*MG1655 +O<sub>2</sub> +aTc (glass tube) | ||
| + | #*MG1655 -O<sub>2</sub> +aTc (15 mL conical tube) | ||
| + | #*MG1655 +CRISPRi +O<sub>2</sub> -aTc (glass tube) | ||
| + | #*MG1655 +CRISPRi -O<sub>2</sub> -aTc (15 mL conical tube) | ||
| + | #*MG1655 +CRISPRi +O<sub>2</sub> +aTc (glass tube) | ||
| + | #*MG1655 +CRISPRi -O<sub>2</sub> +aTc (15 mL conical tube) | ||
| + | #Be sure to include your team information on each tube! | ||
| + | #Obtain a 45 mL aliquot of LB broth from the front laboratory bench. | ||
| + | #Transfer 5 mL of the media into each tube labeled for inoculation with MG1655 (alone, not co-transformed). | ||
| + | #Calculate the volume of the chloramphenicol and ampicillin antibiotic stocks that are required to maintain the pdCas9 and pgRNA plasmids, respectively, in the remaining volume of the LB media using the information below: | ||
| + | #*For chloramphenicol: the stock concentration is 34 mg/mL and the final or 'working' concentration should be 34 μg/mL. | ||
| + | #*For ampicillin: the stock concentration is 100 mg/mL and the final or 'working' concentration should be 100 μg/mL. | ||
| + | #Add the appropriate volume of each antibiotic stock to the LB aliquot and mix thoroughly. | ||
| + | #Transfer 5 mL of the media you prepared in Step #5 into each tube labeled for inoculation with MG1655 +CRISPRi. | ||
| + | On either Monday or Tuesday afternoon, depending on your laboratory section, the teaching faculty will inoculate MG1655 or a co-transformant into your culture tubes and add 2 μM aTc to the appropriate tubes. All tubes will then be incubated at 37 °C until you return for the next laboratory session. | ||
==Reagents== | ==Reagents== | ||
| + | *LB broth | ||
| + | **Luria-Bertani (LB) broth contains 1% tryptone, 0.5% yeast extract, and 1% NaCl | ||
| + | *chloramphenicol antibiotic stock (34 mg/mL) | ||
| + | *ampicillin antibiotic stock (100 mg/mL) | ||
| + | *anhydrotetracycline (aTc, 2 μM, Sigma Aldrich) | ||
==Navigation links== | ==Navigation links== | ||
| − | Next day: [ | + | Next day: [[20.109(F16)::Measure fermentation products (Day8) | Measure fermentation products]] |
Previous day: [[20.109(F16):Journal club II (Day 6)| Journal club II]] | Previous day: [[20.109(F16):Journal club II (Day 6)| Journal club II]] | ||
Latest revision as of 19:38, 3 November 2016
Contents
Introduction
The CRISPRi system involves three genetic elements: the target gene within the host genome, the pdCas9 plasmid, and the pgRNA_target plasmid. Though the target genome is native to the host cell, the plasmids must be transformed into the cell and maintained using antibiotic selection. Throughout this module, we have learned about and worked with the CRISPRi plasmids as individual units, but now we will consider the system as a whole in the context of gene regulation.
In the previous laboratory session, you co-transformed pdCas9 and your pgRNA_target plasmids into E. coli MG1655. The colonies present on your LB plates containing chloramphenicol and ampicillin should carry both plasmids in that they were able to survive selection by both antibiotics. The promoter (pJ23119) driving expression of your gRNA sequence in the gRNA_target plasmid is constitutively active. This means that RNAP is not prohibiting from binding and transcription of the gRNA target sequence is therefore not inhibited. Therefore, your gRNA_target is always present in the MG1655 cells and binding to the target sequence within the genome.
In contrast, expression of the gene encoding Cas9 within pdCas9 is regulated by an inducible promoter (pLtetO-1). An inducible promoter is 'off' unless the appropriate molecule is present to relieve repression. In the case of our system, expression of the gene encoding Cas9 is inhibited due to the use of a tet-based promoter construct. Tet is shorthand for tetracycline, which is an antibiotic that inhibits protein synthesis through preventing the association between charged aminoacyl-tRNA molecules and the A site of ribosomes. Bacterial cells that carry the tet resistance cassette are able to survive exposure to tetracycline by expressing genes that encode an efflux pump that 'flushes' the antibiotic from the bacterial cell. To conserve energy, the tet system is only expressed in the presence of tetracyline. In the absence of tetracycline, a transcription repressor protein (TetR) is bound to the promoter upstream of the tet resistance cassette genes. When tetracycline is present, the molecule binds to TetR causing a confirmational resulting in TetR 'falling off' of the promoter. In the CRISPRi system, the tet-based promoter construct upstream of the gene that encodes Cas9 is 'off' unless anhydrotetracyline (aTc), an analog of tetracyline, is added to the culture media.Taken together, the gRNA-target molecule is constitutively transcribed and, as stated above, always present / binding to the target. The dCas9 protein is only present when aTc is added. Thus, gene expression is only altered when aTc is present. As represented by the schematic below, the gRNA_target 'seeks out' the target within the host genome and recruits dCas9 to the site. When associated with the target / gRNA_target complex, dCas9 binds to the site and acts as a 'roadblock' by prohibiting RNAP access to the sequence. Because the gene of interest is not able to be transcribed, the protein encoded by that gene is not synthesized. In our experiments, we hypothesize that the absence of specific proteins, or enzymes, in the fermentation pathway will increase the yield of either ethanol or lactate.
Protocols
Part 1: Communication Lab workshop
Our communication instructor Dr. Sean Clarke will join us today for a workshop on organizing and writing your M2 research article.
Part 2: Examine pgRNA sequencing results
Your goal today is to analyze the sequencing data for you two potential mutant pgRNA clones - two independent colonies from your amplification reaction - and then decide which colony to proceed with for the CRISPRi manipulation of the E. coli MG1655 fermentation pathway.
Retrieve sequence results from Genewiz
- Use the pgRNA_target Benchling file that you generated on M2D5 to confirm that the gRNA sequence you designed was indeed inserted into the vector.
- Your sequencing data is available from Genewiz. It was uploaded to the Module 2 main page Discussion tab.
You should align your sequencing data with a known sequence, in this case the gRNA target sequence you selected, to identify any unintended base changes that may have occurred. There are several web-based programs for aligning sequences and still more programs that can be purchased. The steps for using Benchling are below. Please feel free to use any program with which you are familiar.
Align with A Plasmid Editor
You can use any DNA manipulation software you choose to complete the alignment confirmation, but the instructions provided are for APE (A Plasmid Editor, created by M. Wayne Davis at the University of Utah). The software can be downloaded free-of-charge from this site onto your personal computer or you can use the 20.109 laboratory computers. Please note that if you use a different program the teaching faculty may not be able to assist you.
- Make an APE DNA file that contains the gRNA oligo you used on M2D3. This sequence should contain the target sequence you selected and the dCas9 tag sequence.
- Open APE then copy and paste the sequence into a new workspace.
- Open a new window and create an APE DNA file that contains the results from your sequencing reaction.
- Go to File and select 'New' to open a new window.
- Paste the sequence text from your sequencing run into the new window. If there were ambiguous areas of your sequencing results, these will be listed as "N" rather than "A" "T" "G" or "C" and it's fine to include Ns in the query.
- The start and end of your sequencing may have several Ns. In this case it is best to omit these Ns by pasting only the 'good' sequence that is flanked by the ambiguous sequence.
- Go to Tools and select 'Align Two Sequences...'
- In one drop-down window choose the gRNA oligo file and in the second drop-down window choose the new file that contains the Genewiz sequence you copied and pasted in step #3.
- Be sure to consider whether you want to compare the reverse-complement of the Genewiz sequence and, if appropriate, check the box to the right of the drop-down window. If you are unsure if this box should be checked, ask the teaching faculty.
- Click 'OK' and a new window should open with the sequences aligned. Matches will be shown by vertical lines between the aligned sequences. You should see a long stream of matches. If your insertion is present, then in this stream of matches the inserted basepairs should be highlighted in red.
- Carefully examine the sequence to see if your gRNA insertion was incorporated.
- You should save a screenshot of each alignment and attach them to your Benchling notebook.
- Follow the above steps to examine all of your sequencing results. Remember: you used a forward and a reverse primer to interrogate both potential gRNA_target plasmids.
If both clones for your gRNA_target have the correct sequence, choose either co-transformant to use for the aTc induction step. If only one is correct, then this is the co-transformant you will use next time. If neither of your plasmids carry the appropriate insertion, talk to the teaching faculty.
Part 3: Prepare media for mixed-acid fermentation inoculations
To test the effect of your gRNA on altering the production of either ethanol or lactate, you will incubate the co-transformed E. coli cells both with and without oxygen. Remember from lecture that cells grown anaerobically will produce more fermentation products; however, our goal is to further enhance the production of these products by manipulating the fermentation pathway.
- Acquire 4 glass culture tubes and 4 15 mL conical tubes from the front laboratory bench and label as shown below:
- MG1655 +O2 -aTc (glass tube)
- MG1655 -O2 -aTc (15 mL conical tube)
- MG1655 +O2 +aTc (glass tube)
- MG1655 -O2 +aTc (15 mL conical tube)
- MG1655 +CRISPRi +O2 -aTc (glass tube)
- MG1655 +CRISPRi -O2 -aTc (15 mL conical tube)
- MG1655 +CRISPRi +O2 +aTc (glass tube)
- MG1655 +CRISPRi -O2 +aTc (15 mL conical tube)
- Be sure to include your team information on each tube!
- Obtain a 45 mL aliquot of LB broth from the front laboratory bench.
- Transfer 5 mL of the media into each tube labeled for inoculation with MG1655 (alone, not co-transformed).
- Calculate the volume of the chloramphenicol and ampicillin antibiotic stocks that are required to maintain the pdCas9 and pgRNA plasmids, respectively, in the remaining volume of the LB media using the information below:
- For chloramphenicol: the stock concentration is 34 mg/mL and the final or 'working' concentration should be 34 μg/mL.
- For ampicillin: the stock concentration is 100 mg/mL and the final or 'working' concentration should be 100 μg/mL.
- Add the appropriate volume of each antibiotic stock to the LB aliquot and mix thoroughly.
- Transfer 5 mL of the media you prepared in Step #5 into each tube labeled for inoculation with MG1655 +CRISPRi.
On either Monday or Tuesday afternoon, depending on your laboratory section, the teaching faculty will inoculate MG1655 or a co-transformant into your culture tubes and add 2 μM aTc to the appropriate tubes. All tubes will then be incubated at 37 °C until you return for the next laboratory session.
Reagents
- LB broth
- Luria-Bertani (LB) broth contains 1% tryptone, 0.5% yeast extract, and 1% NaCl
- chloramphenicol antibiotic stock (34 mg/mL)
- ampicillin antibiotic stock (100 mg/mL)
- anhydrotetracycline (aTc, 2 μM, Sigma Aldrich)
Next day: Measure fermentation products
Previous day: Journal club II