Difference between revisions of "20.109(S16):Module 1"
MAXINE JONAS (Talk | contribs) (→Overview) |
Noreen Lyell (Talk | contribs) (→Overview) |
||
| Line 15: | Line 15: | ||
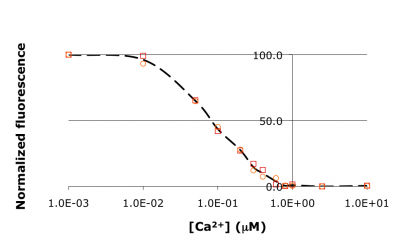
[[Image:20109_S12M2_titration-ex.png|thumb|right|400px|'''Raw titration curve for IPC.''' Shown here is sample data from the teaching lab: normalized fluorescence for wild-type inverse pericam as a function of calcium concentration. As you will later learn, an apparent ''K<sub>d</sub>'' can be estimated from such a plot: it is the point on the ''x''-axis where the curve crosses ''y'' = 50%, or ~0.1 μM here.]] | [[Image:20109_S12M2_titration-ex.png|thumb|right|400px|'''Raw titration curve for IPC.''' Shown here is sample data from the teaching lab: normalized fluorescence for wild-type inverse pericam as a function of calcium concentration. As you will later learn, an apparent ''K<sub>d</sub>'' can be estimated from such a plot: it is the point on the ''x''-axis where the curve crosses ''y'' = 50%, or ~0.1 μM here.]] | ||
| − | + | Your goal in this module is to engineer a protein, inverse pericam (IPC), which was developed by [http://www.ncbi.nlm.nih.gov/pubmed/11248055 Nagai et al.]. IPC is comprised of a permuted enhanced yellow fluorescent protein (EYFP) linked to a calcium sensor. The “inverse” in the name refers to the fact that this protein shines brightly in the absence of calcium, but dimly when calcium is added at a sufficient concentration. You will mutate a single residue within IPC in an attempt to alter the calcium binding properties of the protein. Because IPC is used as a calcium sensor, changing the properties of this protein may generate a novel tool that can be used in currently ill-understood environments or systems. | |
| + | |||
| + | The dissociation constant ''K<sub>d</sub>'' of wild-type IPC with respect to calcium is reported to be 0.2 μM (see figure below). Your goal is to alter the binding curve by mutating a single residue in IPC. You will modify inverse pericam at the gene level using a process called site-directed mutagenesis, express the resultant protein in a bacterial host, and finally purify your mutant protein and assay its calcium-binding activity via a fluorescence assay. You may find that you shift the titration curve to the left or right, which corresponds to altering ''K<sub>d</sub>''; or you might change the steepness of the titration curve, which corresponds to changing cooperativity (and thus concentration dynamic range); finally, you might affect the maximum and/or minimum fluorescence values, thus changing the sensor's signal:noise profile (fluorescence dynamic range). You might even obliterate the response to calcium entirely! In the course of this module, we will consider the benefits and drawbacks of different approaches to protein design, and the types of scientific investigations and applications enabled by fluorescently tagged biological molecules. | ||
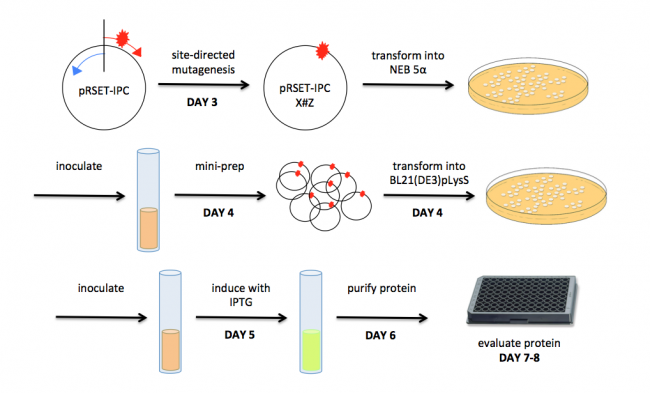
[[Image:Sp16 M1 overview.png|thumb|center|650px|'''Schematic of Module 1.''']] | [[Image:Sp16 M1 overview.png|thumb|center|650px|'''Schematic of Module 1.''']] | ||
Revision as of 17:22, 27 January 2016
Module 1
Lecturer: Noreen Lyell
Instructors: Leslie McClain and Maxine Jonas
TA: Jing Zhang
Lab manager: Hsinhwa Lee
Overview
Your goal in this module is to engineer a protein, inverse pericam (IPC), which was developed by Nagai et al.. IPC is comprised of a permuted enhanced yellow fluorescent protein (EYFP) linked to a calcium sensor. The “inverse” in the name refers to the fact that this protein shines brightly in the absence of calcium, but dimly when calcium is added at a sufficient concentration. You will mutate a single residue within IPC in an attempt to alter the calcium binding properties of the protein. Because IPC is used as a calcium sensor, changing the properties of this protein may generate a novel tool that can be used in currently ill-understood environments or systems.
The dissociation constant Kd of wild-type IPC with respect to calcium is reported to be 0.2 μM (see figure below). Your goal is to alter the binding curve by mutating a single residue in IPC. You will modify inverse pericam at the gene level using a process called site-directed mutagenesis, express the resultant protein in a bacterial host, and finally purify your mutant protein and assay its calcium-binding activity via a fluorescence assay. You may find that you shift the titration curve to the left or right, which corresponds to altering Kd; or you might change the steepness of the titration curve, which corresponds to changing cooperativity (and thus concentration dynamic range); finally, you might affect the maximum and/or minimum fluorescence values, thus changing the sensor's signal:noise profile (fluorescence dynamic range). You might even obliterate the response to calcium entirely! In the course of this module, we will consider the benefits and drawbacks of different approaches to protein design, and the types of scientific investigations and applications enabled by fluorescently tagged biological molecules.
We gratefully acknowledge 20.109 instructors Natalie Kuldell and Prof. Alan Jasanoff for helpful discussions during the development of this module, as well as for their prior work in developing a related module.
Lab links: day by day
M1D1: In silico cloning
M1D2: Design mutation primers
M1D3: Site-directed mutagenesis
M1D4: Prepare expression system
M1D5: Induce protein and evaluate DNA
M1D6: Purify protein
M1D7: Characterize protein expression
M1D8: Assess protein function