Difference between revisions of "20.109:Module 1"
MAXINE JONAS (Talk | contribs) m (13 revisions: Test transfer 20.109(S07) to HostGator) |
|||
| (9 intermediate revisions by 2 users not shown) | |||
| Line 1: | Line 1: | ||
| − | <div style="padding: 10px; width: | + | {{Template:20.109}} |
| + | |||
| + | <div style="padding: 10px; width: 640px; border: 5px solid #2171B7;"> | ||
==Module 1== | ==Module 1== | ||
| − | '''Instructors:''' [ | + | '''Instructors:''' [http://openwetware.org/wiki/Drew_Endy ] and [http://openwetware.org/wiki/Natalie_Kuldell ] |
| − | '''TA:''' [ | + | '''TA:''' [http://openwetware.org/wiki/Sean_Clarke ] |
| + | In this experiment, we will consider the genome of a virus, namely the bacteriophage M13. M13 is a self-assembling nano-machine with a compact genome that has been optimized by evolution to commandeer its bacterial host. Approximately 1000 new viruses are generated from a single infection event. Imagine harnessing this production. What could we build and what natural processes could we better understand? One approach we���ll take is to modify the existing genome in a subtle but useful way, namely by adding a peptide-tag that can be presented on the bacteriophage coat. We���ll examine how this modification affects the coat protein���s expression and overall phage production. Another approach we���ll take is to start from scratch, undertaking a full throttle redesign of the bacteriophage genome. We���ll employ a commercial DNA synthesis company to compile the redesigned genomic program and then we���ll see if it encoded infective M13 and if the genome of the bacterial host affects bacteriophage production. Through these investigations we���ll ask: is nature���s M13 genome ���perfect��� or can we do better? | ||
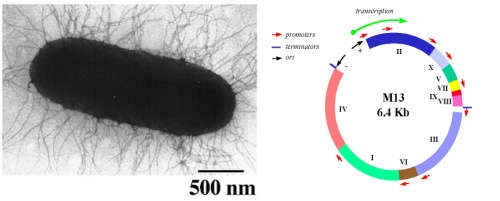
[[Image:Macintosh HD-Users-nkuldell-Desktop-GnmEng coverart S07.jpg|thumb|500px|center|M13-coated coli from M. Russel<br> Map of M13 genome from M. Blaber]] | [[Image:Macintosh HD-Users-nkuldell-Desktop-GnmEng coverart S07.jpg|thumb|500px|center|M13-coated coli from M. Russel<br> Map of M13 genome from M. Blaber]] | ||
| + | |||
| + | [[20.109(S07):Start-up genome engineering | Module 1 Day 1: Start-up genome engineering]]<br> | ||
| + | [[20.109(S07): Agarose gel electrophoresis| Module 1 Day 2: Agarose gel electrophoresis]]<br> | ||
| + | [http://openwetware.org/wiki/20.109(S07):_DNA_ligation_and_bacterial_transformation | Module 1 Day 3: DNA ligation and bacterial transformation]<br> | ||
| + | [http://openwetware.org/wiki/20.109(S07):_Examine_candidate_clones | Module 1 Day 4: Examine candidate clones]<br> | ||
| + | [[20.109(S07): Western analysis| Module 1 Day 5: Western analysis]]<br> | ||
| + | [http://openwetware.org/wiki/20.109(S07):_Probe_western | Module 1 Day 6: Probe western]<br> | ||
| + | direct link to [[20.109%28S07%29:_Genome_engineering_essay]]<br> | ||
| + | direct link to [http://openwetware.org/wiki/20.109:Module_1:RefactorM13 | M13 refactoring workpage]<br> | ||
| + | direct link to [http://parts.mit.edu/r/parts/partsdb/part_info.cgi?part_name=BBa_M1307| hard info for BBa_M1307] | ||
| + | |||
| + | [http://openwetware.org/wiki/20.109(S07):_TA's_notes_for_module_1 | TA notes, mod 1] | ||
Latest revision as of 15:31, 15 June 2015
Module 1
TA: [3]
In this experiment, we will consider the genome of a virus, namely the bacteriophage M13. M13 is a self-assembling nano-machine with a compact genome that has been optimized by evolution to commandeer its bacterial host. Approximately 1000 new viruses are generated from a single infection event. Imagine harnessing this production. What could we build and what natural processes could we better understand? One approach we���ll take is to modify the existing genome in a subtle but useful way, namely by adding a peptide-tag that can be presented on the bacteriophage coat. We���ll examine how this modification affects the coat protein���s expression and overall phage production. Another approach we���ll take is to start from scratch, undertaking a full throttle redesign of the bacteriophage genome. We���ll employ a commercial DNA synthesis company to compile the redesigned genomic program and then we���ll see if it encoded infective M13 and if the genome of the bacterial host affects bacteriophage production. Through these investigations we���ll ask: is nature���s M13 genome ���perfect��� or can we do better?
Module 1 Day 1: Start-up genome engineering
Module 1 Day 2: Agarose gel electrophoresis
| Module 1 Day 3: DNA ligation and bacterial transformation
| Module 1 Day 4: Examine candidate clones
Module 1 Day 5: Western analysis
| Module 1 Day 6: Probe western
direct link to 20.109(S07):_Genome_engineering_essay
direct link to | M13 refactoring workpage
direct link to hard info for BBa_M1307