Difference between revisions of "20.109(S08):Site-directed mutagenesis (Day2)"
(→Protocols) |
MAXINE JONAS (Talk | contribs) m (92 revisions) |
||
| (76 intermediate revisions by 2 users not shown) | |||
| Line 1: | Line 1: | ||
{{Template:20.109(S08)}} | {{Template:20.109(S08)}} | ||
| + | |||
| + | |||
| + | ''First, check out some general [http://openwetware.org/wiki/20.109(S08):_Module_2_General_Comments_on_Lab_Notebooks |comments] on your lab notebooks. | ||
| + | '' | ||
==Introduction== | ==Introduction== | ||
| + | |||
| + | Last time you navigated a great deal of information in order to design mutagenized inverse pericams ��� nice work! Today you will put your designs into practice. | ||
| + | |||
| + | [[Image:20.109_SDM-Nobel.png|thumb|right|150px| 1993 Chemistry Nobel Prize co-winner (with Kary Mullis, inventor of PCR) for developing site-directed mutagenesis.]] | ||
| + | As you already know, the mutagenesis strategy you will use shares some features with the PCR you performed in Module 1. You will begin by combining plasmid DNA encoding wild-type inverse pericam with the mutagenic primers you designed. These will be acted upon by a DNA polymerase to generate a mutant plasmid ��� at this stage the plasmid will be nicked, and won���t become whole until it is transformed into competent bacteria and repaired by them. In order to propagate only the mutant plasmid, the parental DNA is specifically digested using the DpnI enzyme prior to bacterial transformation. The thermocycling reaction will run for about two hours, and the digestion step for another hour. During these incubation times, we will discuss two articles from the primary literature. | ||
| + | |||
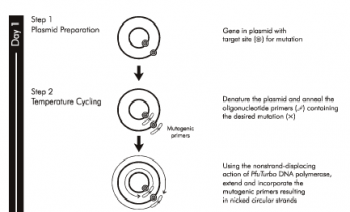
| + | [[Image:20.109_SG-SDM-P1.png|thumb|left|350px| '''From Stratagene QuikChange® Manual, Part 1''']] | ||
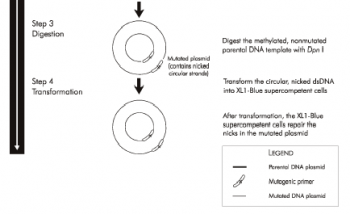
| + | [[Image:20.109_SG-SDM-P2.png|thumb|none|350px| '''From Stratagene QuikChange® Manual, Part 2''']] | ||
| + | |||
| + | <br style="clear:both;"/> | ||
| + | |||
| + | We will ���warm-up��� by discussing the paper by [http://www.ncbi.nlm.nih.gov/pubmed/7809066?ordinalpos=9&itool=EntrezSystem2.PEntrez.Pubmed.Pubmed_ResultsPanel.Pubmed_RVDocSum Heim, Prasher, and Tsien]. This is a short paper describing the very first attempt to mutagenize GFP, and a fine introduction to some of the concepts and methods used in this module. Next we will do a close reading of the paper that introduced inverse pericam, by [http://www.ncbi.nlm.nih.gov/pubmed/11248055?ordinalpos=5&itool=EntrezSystem2.PEntrez.Pubmed.Pubmed_ResultsPanel.Pubmed_RVDocSum Nagai et al]. We will examine the construction and analysis of the IPC multi-component calcium sensor in some depth. | ||
| + | |||
| + | Now might be a good time to mention why we care about measuring intracellular calcium in the first place. Calcium is involved in many signal transduction cascades, which regulate everything from immune cell activation to muscle contraction, from adhesion to apoptosis - see for example | ||
| + | [http://www.ncbi.nlm.nih.gov/pubmed/18083096?ordinalpos=1&itool=EntrezSystem2.PEntrez.Pubmed.Pubmed_ResultsPanel.Pubmed_RVDocSum this review by David Clapham in Cell], or [http://www.ncbi.nlm.nih.gov/pubmed/11830654?ordinalpos=6&itool=EntrezSystem2.PEntrez.Pubmed.Pubmed_ResultsPanel.Pubmed_RVDocSum this one by Ernesto Carafoli in PNAS]. Intracellular calcium (Ca<sup>2+</sup>) is normally maintained at ~100 nM, orders of magnitude less than the ~mM concentration outside the cell. ATPase pumps act to keep the basal concentration of cytoplasmic calcium low. Often calcium acts as a secondary messenger, i.e., it relays a message from the cell surface to its cytoplasm. For example, a particular ligand may bind a cell surface receptor, causing a flood of calcium ions to be released from the intracellular compartments in which they are usually sequestered. These free ions in turn may promote phosphorylation or other downstream signaling. | ||
| + | |||
| + | The proteins that bind calcium do so with a great variety of affinities, and have roles ranging from sequestration to sensing. Some calcium responses may have long-term effects, particularly in the case of transcription factors that can bind calcium. As you learned last time, calmodulin works as a calcium sensor by undergoing a conformational change upon calcium binding. Your goal today is to prepare mutant calmodulin (in the context of inverse pericam) DNA, in order to alter the affinity of the resulting protein for calcium. | ||
==Protocols== | ==Protocols== | ||
| − | ===Part 1: | + | ===Part 1: Primer preparation=== |
| − | #Calculate the amount of water needed ''for each primer'' to give a concentration of 1 | + | #Calculate the amount of water needed ''for each primer'' to give a concentration of 1 mg/mL. |
#Touch-spin your primers, resuspend each in the appropriate volume of water, and touch-spin again. | #Touch-spin your primers, resuspend each in the appropriate volume of water, and touch-spin again. | ||
| − | #Calculate the dilution of primer that you need to prepare in order to have 125 ng present in 2.5 | + | #Calculate the dilution of primer that you need to prepare in order to have 125 ng present in 2.5 ��L. Prepare 100-200 ��L of this dilution and keep it on ice. |
| − | ===Part 2: | + | ===Part 2: Site-directed mutagenesis=== |
| − | We will be using the | + | We will be using the QuickChange�� kit from Stratagene to perform our site-directed mutageneses. Each group will set up two reactions. Meanwhile, your TA will set up a single positive control reaction, to ensure that all the reagents are working properly. You should work quickly but carefully, and keep your tubes in a chilled container at all times. |
#Read through the following protocol and prepare all calculations before beginning physical manipulations of your samples. | #Read through the following protocol and prepare all calculations before beginning physical manipulations of your samples. | ||
| − | #You will be given | + | #You will be given a 95 ��L mixture that contains buffer and dNTPs when you are ready to begin. Aliquot 43 ��L per PCR tube. |
| − | #Add 2 | + | #Add 2 ��L of template DNA (���IPC plasmid���) to each of your two reaction tubes. |
| − | #*Note: mutagenesis reactions are expected to run smoothly with 5-50 ng of plasmid DNA. You have been given a 1:200 dilution of miniprep DNA. | + | #*Note: mutagenesis reactions are expected to run smoothly with 5-50 ng of plasmid DNA. You have been given a 1:200 dilution of miniprep DNA. |
| − | # | + | #Per mutation reaction, add 2.5 μL of each primer (forward and reverse) to the right tube. Do not mix all four of your primers together! |
| − | #Finally, add 1 | + | #The volume of each reaction should be now be 50 ��L. |
| + | #Finally, add 1 ��L of ''PfuTurbo'' DNA polymerase to each reaction. | ||
#Once each group is ready, we will begin the thermocycler, under the following conditions: | #Once each group is ready, we will begin the thermocycler, under the following conditions: | ||
| Line 59: | Line 81: | ||
| − | #After the cycling is completed, each group should add 1 | + | #After the cycling is completed, each group should add 1 ��L of ''DpnI'' per tube. Samples will be incubated for one hour at 37 ° C. You will be asked to how this enzyme works in your homework assignments. |
| − | #This final incubation step will take us past lab closing time, at which point the teaching staff will take over. Each reaction will be transformed into XL1-Blue competent cells under delicate conditions. | + | #This final incubation step will take us past lab closing time, at which point the teaching staff will take over. Each reaction will be transformed into XL1-Blue competent cells under delicate conditions.(*) |
#Tomorrow, two candidate colonies will be chosen from each plate. The efficiency of this mutagenesis protocol is reported to be 80%. We will sequence two candidates per mutation to cover our bases, so to speak. | #Tomorrow, two candidate colonies will be chosen from each plate. The efficiency of this mutagenesis protocol is reported to be 80%. We will sequence two candidates per mutation to cover our bases, so to speak. | ||
| + | |||
| + | (*)<small>In a 14 mL polypropylene round-bottom tube, 1 μL of DNA will be added to 50 μL of competent cells, and the mixture kept on ice for 10 minutes. The cells will be placed in a 42 °C water bath for exactly 45 seconds, back on ice for 2 minutes, and then in a 37 °C incubator for 30 minutes after the addition of 0.5 mL of ''pre-warmed'' LB medium. Finally, 250 μL of cells will be plated on LB-Amp plates as usual.</small> | ||
===Part 3: Prepare tubes for liquid O/N cultures=== | ===Part 3: Prepare tubes for liquid O/N cultures=== | ||
| − | You will make your teaching faculty very happy if you contribute to their preparatory work. Please label 4 large glass test tubes with your team color and sample name. Mix 12 ml LB with 12 μL | + | You will make your teaching faculty very happy if you contribute to their preparatory work. Please label 4 large glass test tubes with your team color and sample name. Mix 12 ml LB with 12 μL of ampicillin. Aliquot 2.5 mL of LB+Amp per tube. These will be used to set up overnight cultures for you for next time. |
| − | ===Part 4: | + | ===Part 4: Journal article discussion=== |
| − | During the production of the mutagenized DNA, we will discuss the | + | During the production of the mutagenized DNA, we will discuss the two journal articles cited in the introduction. The purpose of this discussion will be two-fold: 1) to familiarize ourselves with the history of protein design, and 2) to explore ways of talking about the scientific literature. |
| + | |||
| + | As you read the paper by [http://www.ncbi.nlm.nih.gov/pubmed/7809066?ordinalpos=9&itool=EntrezSystem2.PEntrez.Pubmed.Pubmed_ResultsPanel.Pubmed_RVDocSum Heim, Prasher, and Tsien], consider the following questions. | ||
| + | |||
| + | *What are the advantages of GFP compared to synthetic fluorescent dyes, and what are its limitations? Which of these limitations are Heim et al. trying to address?<br> | ||
| + | *What methods did the authors use for mutagenesis, and how do they compare to the method we are using? <br> | ||
| + | *How were mutagenic proteins initially selected, and how were the chosen ones further analyzed? <br> | ||
| + | *What is the significance of the different wild-type protein fractions shown in Figure 1? <br> | ||
| + | *If GFP maturation does not require any cofactors for a chemical reaction, why does it take four hours? What lines of evidence suggest the absence of cofactors? <br> | ||
| + | *In what part(s) of the protein were useful amino acid substitutions found? <br> | ||
| + | |||
| + | When you arrive in lab today, each group will be assigned one of the following numbered topics to present to and discuss with the rest of the class. You should be somewhat familiar with the whole [http://www.ncbi.nlm.nih.gov/pubmed/11248055?ordinalpos=5&itool=EntrezSystem2.PEntrez.Pubmed.Pubmed_ResultsPanel.Pubmed_RVDocSum Nagai et al] paper by now, but will have some time in-class to refresh your memory and become the resident expert in one of the following areas: | ||
| + | |||
| + | #Introduction and Figure 1 | ||
| + | #*Consider: What is a chimeric protein? A circularly permuted one? | ||
| + | #*How does the chimeric protein pericam work as a sensor? | ||
| + | #Figure 2 and wrap-up at the end | ||
| + | #*What factors - besides the mutations shown in Table 1 ��� affected the fluorescence properites of the pericams? | ||
| + | #*How does the wavelength selected by the authors for testing pericam compare to the excitation maximum of cpEYFP? | ||
| + | #*(wrap-up) Describe some other calcium indicators, and how they compare to pericam. How does the concept of FRET come into play? | ||
| + | #Figure 3 and Table 1 | ||
| + | #*Describe the three types of pericam initially constructed and tested. | ||
| + | #*What kinds of amino acid substitutions were made, and why might they cause the noted effects? | ||
| + | #Figure 4 | ||
| + | #*What method was used to change from bacterial to mammalian expression of pericam? | ||
| + | #*What chemicals can be used to create step-changes in intracellular calcium, and how do some of them work? | ||
| + | #Figure 5 | ||
| + | #*How did the authors calibrate calcium levels? | ||
| + | #*What���s special about the way split-pericam functions? | ||
| + | #Figure 6 | ||
| + | #*How does pericam improve upon previous limitations to imaging calcium in specific cellular compartments? | ||
| + | #*What were the authors able to learn about calcium transients in different organelles? | ||
| + | |||
| + | Finally, you should all consider the similarities and differences between the research described in the papers above and the research that you are undertaking in this module. | ||
==For next time== | ==For next time== | ||
| + | |||
| + | #Calculate how much plasmid DNA you used per mutagenesis reaction, assuming a typical miniprep concentration of 1 μg/μL. | ||
| + | #The major assessment for this module will be a three-part [[20.109%28S08%29:Module_2_Portfolio |portfolio]]. Part 1 asks you to describe your design process and ultimately the two design choices you made for mutant inverse pericams. Get a head start on this assignment by structuring what you want to talk about in outline form (e.g., paragraph 1 = introduction to inverse pericam, paragraph 2 = options for mutations considered, etc.). For each paragraph heading, write a few talking points that you will expand on later (e.g., for paragraph 1: two main components; how calcium sensor works; how linked to fluorescence molecule; why useful). You can write a few sentences if it helps you better organize your thoughts, but short phrases are perfectly fine. You will be graded only on structure and high-level content, not at the word and sentence level. | ||
| + | #Part 2 of your portfolio will be a summary of your data, similar to a combined Results+Discussion section of a report. Knock one figure out of the way this weekend: make a schematic that shows the two mutations you made, indicating both the nucleotide and amino acid level changes, and write a short caption for it. | ||
| + | #Let���s take a moment to explore the smallest component of the inverse pericam construct. What protein is the M13 peptide derived from, and what does it usually do? | ||
| + | |||
| + | ==Reagent List== | ||
| + | |||
| + | *Site-Directed Mutagenesis Kit from Stratagene | ||
| + | **10X Reaction Buffer (100 mM KCl, 100 mM (NH<sub>4</sub>)<sub>2</sub>SO<sub>4</sub>, 200 mM Tris-HCl, 20 mM MgSO<sub>4</sub>, 1% Triton® X-100, 1 mg/mL BSA) | ||
| + | **''PfuTurbo''® DNA polymerase (2.5 U/μL, 1 μL per rxn) | ||
| + | **dNTP mix (proprietary mix, 1 μL per rxn) | ||
| + | **''Dpn I'' (10 U/μL) | ||
| + | **XL1-Blue supercompetent cells | ||
| + | |||
| + | *LB (Luria-Bertani broth) | ||
| + | **1% Tryptone | ||
| + | **0.5% Yeast Extract | ||
| + | **1% NaCl | ||
| + | ** autoclaved for sterility | ||
| + | |||
| + | *Ampicillin: 100 mg/mL, aqueous, sterile-filtered | ||
| + | |||
| + | *LB+AMP plates | ||
| + | **LB with 2% agar and 100 μg/ml Ampicillin | ||
Latest revision as of 14:24, 5 June 2015
First, check out some general |comments on your lab notebooks.
Contents
Introduction
Last time you navigated a great deal of information in order to design mutagenized inverse pericams ��� nice work! Today you will put your designs into practice.
As you already know, the mutagenesis strategy you will use shares some features with the PCR you performed in Module 1. You will begin by combining plasmid DNA encoding wild-type inverse pericam with the mutagenic primers you designed. These will be acted upon by a DNA polymerase to generate a mutant plasmid ��� at this stage the plasmid will be nicked, and won���t become whole until it is transformed into competent bacteria and repaired by them. In order to propagate only the mutant plasmid, the parental DNA is specifically digested using the DpnI enzyme prior to bacterial transformation. The thermocycling reaction will run for about two hours, and the digestion step for another hour. During these incubation times, we will discuss two articles from the primary literature.
We will ���warm-up��� by discussing the paper by Heim, Prasher, and Tsien. This is a short paper describing the very first attempt to mutagenize GFP, and a fine introduction to some of the concepts and methods used in this module. Next we will do a close reading of the paper that introduced inverse pericam, by Nagai et al. We will examine the construction and analysis of the IPC multi-component calcium sensor in some depth.
Now might be a good time to mention why we care about measuring intracellular calcium in the first place. Calcium is involved in many signal transduction cascades, which regulate everything from immune cell activation to muscle contraction, from adhesion to apoptosis - see for example this review by David Clapham in Cell, or this one by Ernesto Carafoli in PNAS. Intracellular calcium (Ca2+) is normally maintained at ~100 nM, orders of magnitude less than the ~mM concentration outside the cell. ATPase pumps act to keep the basal concentration of cytoplasmic calcium low. Often calcium acts as a secondary messenger, i.e., it relays a message from the cell surface to its cytoplasm. For example, a particular ligand may bind a cell surface receptor, causing a flood of calcium ions to be released from the intracellular compartments in which they are usually sequestered. These free ions in turn may promote phosphorylation or other downstream signaling.
The proteins that bind calcium do so with a great variety of affinities, and have roles ranging from sequestration to sensing. Some calcium responses may have long-term effects, particularly in the case of transcription factors that can bind calcium. As you learned last time, calmodulin works as a calcium sensor by undergoing a conformational change upon calcium binding. Your goal today is to prepare mutant calmodulin (in the context of inverse pericam) DNA, in order to alter the affinity of the resulting protein for calcium.
Protocols
Part 1: Primer preparation
- Calculate the amount of water needed for each primer to give a concentration of 1 mg/mL.
- Touch-spin your primers, resuspend each in the appropriate volume of water, and touch-spin again.
- Calculate the dilution of primer that you need to prepare in order to have 125 ng present in 2.5 ��L. Prepare 100-200 ��L of this dilution and keep it on ice.
Part 2: Site-directed mutagenesis
We will be using the QuickChange�� kit from Stratagene to perform our site-directed mutageneses. Each group will set up two reactions. Meanwhile, your TA will set up a single positive control reaction, to ensure that all the reagents are working properly. You should work quickly but carefully, and keep your tubes in a chilled container at all times.
- Read through the following protocol and prepare all calculations before beginning physical manipulations of your samples.
- You will be given a 95 ��L mixture that contains buffer and dNTPs when you are ready to begin. Aliquot 43 ��L per PCR tube.
- Add 2 ��L of template DNA (���IPC plasmid���) to each of your two reaction tubes.
- Note: mutagenesis reactions are expected to run smoothly with 5-50 ng of plasmid DNA. You have been given a 1:200 dilution of miniprep DNA.
- Per mutation reaction, add 2.5 μL of each primer (forward and reverse) to the right tube. Do not mix all four of your primers together!
- The volume of each reaction should be now be 50 ��L.
- Finally, add 1 ��L of PfuTurbo DNA polymerase to each reaction.
- Once each group is ready, we will begin the thermocycler, under the following conditions:
| Segment | Cycles | Temperature (° C) | Time |
|---|---|---|---|
| 1 | 1 | 95 | 30 sec |
| 2 | 18 | 95 | 30 sec |
| 55 | 1 min | ||
| 68 | 5 min | ||
| 3 | 1 | 4 | indefinite |
- After the cycling is completed, each group should add 1 ��L of DpnI per tube. Samples will be incubated for one hour at 37 ° C. You will be asked to how this enzyme works in your homework assignments.
- This final incubation step will take us past lab closing time, at which point the teaching staff will take over. Each reaction will be transformed into XL1-Blue competent cells under delicate conditions.(*)
- Tomorrow, two candidate colonies will be chosen from each plate. The efficiency of this mutagenesis protocol is reported to be 80%. We will sequence two candidates per mutation to cover our bases, so to speak.
(*)In a 14 mL polypropylene round-bottom tube, 1 μL of DNA will be added to 50 μL of competent cells, and the mixture kept on ice for 10 minutes. The cells will be placed in a 42 °C water bath for exactly 45 seconds, back on ice for 2 minutes, and then in a 37 °C incubator for 30 minutes after the addition of 0.5 mL of pre-warmed LB medium. Finally, 250 μL of cells will be plated on LB-Amp plates as usual.
Part 3: Prepare tubes for liquid O/N cultures
You will make your teaching faculty very happy if you contribute to their preparatory work. Please label 4 large glass test tubes with your team color and sample name. Mix 12 ml LB with 12 μL of ampicillin. Aliquot 2.5 mL of LB+Amp per tube. These will be used to set up overnight cultures for you for next time.
Part 4: Journal article discussion
During the production of the mutagenized DNA, we will discuss the two journal articles cited in the introduction. The purpose of this discussion will be two-fold: 1) to familiarize ourselves with the history of protein design, and 2) to explore ways of talking about the scientific literature.
As you read the paper by Heim, Prasher, and Tsien, consider the following questions.
- What are the advantages of GFP compared to synthetic fluorescent dyes, and what are its limitations? Which of these limitations are Heim et al. trying to address?
- What methods did the authors use for mutagenesis, and how do they compare to the method we are using?
- How were mutagenic proteins initially selected, and how were the chosen ones further analyzed?
- What is the significance of the different wild-type protein fractions shown in Figure 1?
- If GFP maturation does not require any cofactors for a chemical reaction, why does it take four hours? What lines of evidence suggest the absence of cofactors?
- In what part(s) of the protein were useful amino acid substitutions found?
When you arrive in lab today, each group will be assigned one of the following numbered topics to present to and discuss with the rest of the class. You should be somewhat familiar with the whole Nagai et al paper by now, but will have some time in-class to refresh your memory and become the resident expert in one of the following areas:
- Introduction and Figure 1
- Consider: What is a chimeric protein? A circularly permuted one?
- How does the chimeric protein pericam work as a sensor?
- Figure 2 and wrap-up at the end
- What factors - besides the mutations shown in Table 1 ��� affected the fluorescence properites of the pericams?
- How does the wavelength selected by the authors for testing pericam compare to the excitation maximum of cpEYFP?
- (wrap-up) Describe some other calcium indicators, and how they compare to pericam. How does the concept of FRET come into play?
- Figure 3 and Table 1
- Describe the three types of pericam initially constructed and tested.
- What kinds of amino acid substitutions were made, and why might they cause the noted effects?
- Figure 4
- What method was used to change from bacterial to mammalian expression of pericam?
- What chemicals can be used to create step-changes in intracellular calcium, and how do some of them work?
- Figure 5
- How did the authors calibrate calcium levels?
- What���s special about the way split-pericam functions?
- Figure 6
- How does pericam improve upon previous limitations to imaging calcium in specific cellular compartments?
- What were the authors able to learn about calcium transients in different organelles?
Finally, you should all consider the similarities and differences between the research described in the papers above and the research that you are undertaking in this module.
For next time
- Calculate how much plasmid DNA you used per mutagenesis reaction, assuming a typical miniprep concentration of 1 μg/μL.
- The major assessment for this module will be a three-part portfolio. Part 1 asks you to describe your design process and ultimately the two design choices you made for mutant inverse pericams. Get a head start on this assignment by structuring what you want to talk about in outline form (e.g., paragraph 1 = introduction to inverse pericam, paragraph 2 = options for mutations considered, etc.). For each paragraph heading, write a few talking points that you will expand on later (e.g., for paragraph 1: two main components; how calcium sensor works; how linked to fluorescence molecule; why useful). You can write a few sentences if it helps you better organize your thoughts, but short phrases are perfectly fine. You will be graded only on structure and high-level content, not at the word and sentence level.
- Part 2 of your portfolio will be a summary of your data, similar to a combined Results+Discussion section of a report. Knock one figure out of the way this weekend: make a schematic that shows the two mutations you made, indicating both the nucleotide and amino acid level changes, and write a short caption for it.
- Let���s take a moment to explore the smallest component of the inverse pericam construct. What protein is the M13 peptide derived from, and what does it usually do?
Reagent List
- Site-Directed Mutagenesis Kit from Stratagene
- 10X Reaction Buffer (100 mM KCl, 100 mM (NH4)2SO4, 200 mM Tris-HCl, 20 mM MgSO4, 1% Triton® X-100, 1 mg/mL BSA)
- PfuTurbo® DNA polymerase (2.5 U/μL, 1 μL per rxn)
- dNTP mix (proprietary mix, 1 μL per rxn)
- Dpn I (10 U/μL)
- XL1-Blue supercompetent cells
- LB (Luria-Bertani broth)
- 1% Tryptone
- 0.5% Yeast Extract
- 1% NaCl
- autoclaved for sterility
- Ampicillin: 100 mg/mL, aqueous, sterile-filtered
- LB+AMP plates
- LB with 2% agar and 100 μg/ml Ampicillin