20.109(S21):Module 1
Contents
Module 1
Lecturer: Dr. Leslie McClain
Instructors: Dr. Noreen Lyell and Dr. Leslie McClain, and Dr. Becky Meyer
Research assistant: Sarah Cowles
TAs: Jeff Hsaio and Caleb Perez
Engineering Antibodies Using Yeast Display
An antibody is a soluble or membrane bound protein, produced by immune cells in blood. In our bodies the purpose of an antibody is to recognize foreign substances, called antigens, and mark them for removal by cells that compromise our immune system. If one has a properly functioning immune system you've all been making new and improved antibodies your entire life. Antibodies are also a critical reagent in medicine, research and diagnostics. Scientists have developed methods to induce and purify antibodies from animals, and also to engineer cells to produce antibodies.
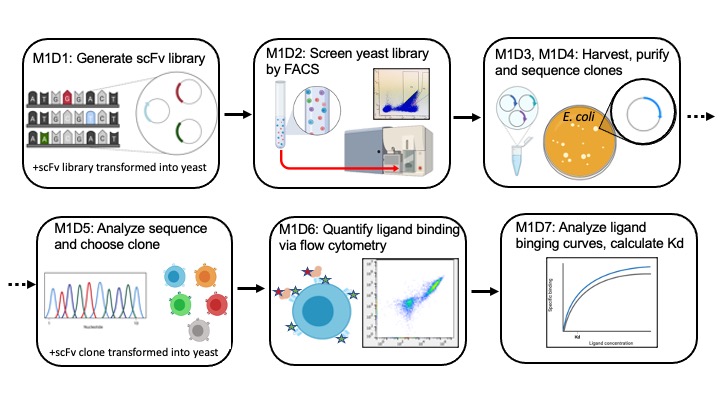
In Mod1 we'll use a method developed in the Wittrup lab to screen a library of antibodies, specifically we will screen a library of antibody fragments, called scFVs or single-chain antibody fragments. These scFvs are expressed on the exterior of yeast cells. We will select for scFVs that have improved binding to an antigen, also called a ligand. In this module we will use lysozyme as our antigen. Lysozyme is a protein that is commercially valuable and used extensively in studies of antibody-antigen binding. The screen will involve the following major steps (1) express scFV library via yeast surface display, (2) incubate scFv yeast with lysozyme, (3) collect fraction of yeast that show strong lysozyme binding via Fluorescence Assisted Cell Sorting (FACS), (4) harvest plasmid DNA from yeast and determine sequence of novel scFv gene (also called clone), (5) characterize novel scFv binding to lysozyme via flow cytometry analysis, (5) calculate binding affinity to determine if binding is improved over parental scFv clone.
This module was developed thanks to the invaluable help and support of Wittrup lab PhD student, Sarah Cowles, and the generous contribution of reagents from Prof. Wittrup.
Research goal: Identify and characterize an antibody fragment (scFv) that shows improved binding to the antigen, lysozyme.
Acronyms and common terms-
scFv: single-chain antibody fragments
clone: novel scFv mutant from library
ligand: molecule that binds to another molecule, interchangeable with the term antigen in this module. Antigen used in this module is the protein lysozyme.
FACS: Fluorescence Assisted Cell Sorting, method to measure physical properties of cells and / or fluorescence on the cells. Cells are kept alive and sorted into distinct tubes to regrow after analysis.
Flow cytometry: method to measure physical properties of cells and / or fluorescence. Cells are moved to biowaste after analysis.
Lab links: day by day
M1D1: Generate scFv library
M1D2: Enrich candidate clones from library using FACS
M1D3: Harvest clone plasmids from library
M1D4: Purify and sequence clone plasmids
M1D5: Analyze clone sequences and choose clone to characterize
M1D6: Characterize clone ligand binding using flow cytometry
M1D7: Analyze ligand titration curves
Major assignments
Data summary
Mini-presentation
References
This module modified from protocols found in this literature:
- Isolating and engineering human antibodies using yeast surface display. Nature Protocols. 1, p755–768(2006).
- Yeast Surface Display for Antibody Isolation: Library Construction, Library Screening, and Affinity Maturation. Methods in Molecular Biology. 1131:151-81(2014).
- Applications of Yeast Surface Display for Protein Engineering. Methods in Molecular Biology (2015) 1319:155-75.
Examples of characterization of scFv clones:
- Yeast Surface Display for Screening Combinatorial Polypeptide Libraries. Nature Biotechnology (1997) 15, 553–557.
- Thermodynamics Characterization of affinity Maturation: the D1.3 antibody and a Higher Affinity Mutant. J. Mol. Recognition (1998) 11, 10-13.
Examples of Using Yeast Surface Display for Non-antibody Proteins:
- Evolution of an Interloop Disulfide Bond in High-Affinity Antibody Mimics Based on FNIII Domainand Selected by Yeast Display. J. Mol. Biol. (2007) 368, 1024–1041.
- Picomolar Affinity Fibronectin Domains Engineered Utilizing Loop Length Diversity Recursive Mutagenesis and Loop Shuffling. J. Mol. Biol. (2008) 381, 1238–1252.
- Beyond Epitope Binning: Directed in Vitro Selection of Complementary Pairs of Binding Proteins. ACS Comb. Sci. (2020) 22, 49−60.
Cellular and Molecular Immunology E-Book available through MIT libraries:
Next steps in antibody development, protocol for expressing full antibodies in mammalian cells:
- Transient expression of human antibodies in mammalian cells. Nature Protocols (2018) 13(10), 99-117