20.109(S17):Assess cell survival and harvest RNA for quantitative PCR assay (Day3)
Contents
Introduction
Differential gene expression is used to translate information perceived by a cell into a gene product. All forms of life rely on gene expression to generate proteins and non-protein coding gene products that provide the machinery for cell survival. In this, gene expression is the most basic mechanism by which genotype results in a phenotype. One method researchers use to examine gene expression is to quantify transcript levels for specific genes.
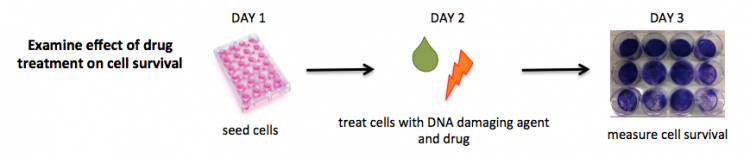
In this module, we are interested in the effect of NHEJ-inhibiting drugs on cell survival and gene expression. To address these questions we are using a BRCA2- mutant cell line. When DNA damage is induced in these cells (either naturally or with the chemical etoposide), the absence of BRCA2 results in a dependence or addiction to the NHEJ pathway. If a drug is introduced following DNA damage that inhibits NHEJ repair, the expected outcome is decreased cell survival compared to wild-type cells; hence the reason for the crystal violet cell survival assay. The more complicated question is how to assess which genes were differentially expressed in response to the drug treatment following DNA damage.
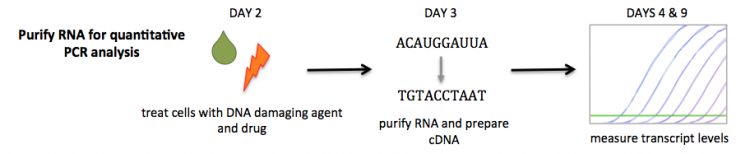
We will use two experimental approaches to evaluate gene expression: qPCR and RNA-seq. Though the intricacies of each method will be discussed on D4 and D7, respectively, a critical first step for both is the purification of high-quality RNA. RNA purification is complicated by ribonuclease enzymes, such as RNase. RNases are ubiquitous can be difficult to neutralize. Furthermore, RNases are are present in the cells and tissues from which the RNA is isolated. The 'gold standard' for RNA purification is an organic extraction method. In this process a sample is homogenized in a phenol solution then centrifuged, which separates the sample into three phases: a lower organic layer, a denatured protein and gDNA middle layer, and an RNA upper layer. The RNA from the upper layer is alcohol precipitated and rehydrated. Because this method is labor-intensive and requires chlorinated organic reagents, researchers often use column-based purification kits.
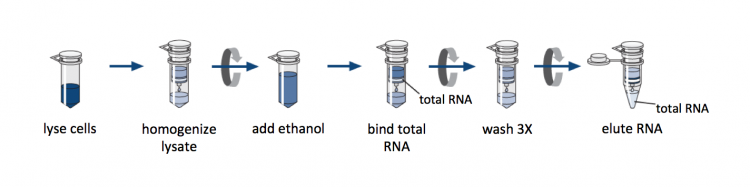
We will use a commercially available column-based RNA purification system. Though the reagents are proprietary, the manufacturer provides some details concerning the inner workings of the kit. The key component is a silica-based membrane that binds RNA molecules longer that 200 nucleotides, which enriches for mRNA molecules. Prior to RNA isolation using the membrane, cells are lysed in the presence of a guanidine-thiocyanate-containing buffer that inactivates the RNases naturally present in the sample. Then ethanol is added to the lysate to promote binding of the RNA to the silica.
Following RNA purification, you will generate cDNA to use in your quantitative PCR analysis. Complementary DNA, or cDNA, refers to DNA that is generated from RNA. The ability to natively generate cDNA is unique to retroviruses. In research, this technology is available as a tool because the reverse transcriptase enzyme can be purified and used in exogenous reactions. We will generate cDNA from purified RNA to study gene expression as DNA molecules are more stable and, therefore, easier to work with in experiments. As with the RNA purification, we will use a kit to generate cDNA. The kit includes the necessary buffers and reverse transriptase enzyme. First, the RNA is combined with a poly-T primer and oligonucleotides, then incubated at a high temperature for a denaturation step. An RNase inhibitor, reverse transcriptase, and buffers are then added. This mix is incubated at a temperature optimal for reverse transcriptase enzyme to generate cDNA. The specific enzyme we will use is SuperScript III RT, and like the buffers included in the RNA purification kit, the genetic manipulations used to modify this reverse transcriptase enzyme are proprietary.
Protocols
Part 1: Workshop with BE Communication Lab
Our communication instructor, Dr. Diana Chien, will join us today for a workshop on transforming a written journal article into a well-defined oral presentation.
Part 2: Prepare samples for quantitative PCR assay
You will complete the first part of this exercise in tissue culture and then move into the main laboratory to purify the RNA from your cells. Before you begin your work in tissue culture, prepare your laboratory bench for work with RNA. As noted in the introduction, RNA is very sensitive to degradation and caution must be taken to preserve your sample.
First, obtain a piece of absorbent paper. This will be your RNAse free work space. Spray the absorbent paper with RNase Away and dry with a Kimwipe. Perform this cleaning procedure with all of the equipment you will need for the RNA purification protocol (read through Part 2a Step #11 to the end of the protocol). Also, obtain all of the aliquots you will need from the front laboratory bench and clean before placing the tubes on the absorbent paper work space. Lastly, obtain a 50 mL conical tube and label it 'waste'.
Part 2a: Purify RNA from cells
- Prepare your working space within the tissue culture hood.
- Retrieve your four T25 flasks (DLD-1 +etop, BRCA2- +etop, DLD-1 +etop +drug, and BRCA2- +etop +drug) from the 37 °C incubator and visually inspect your cells with a microscope.
- Record your observations concerning media color, confluency, etc. in your laboratory notebook.
- Aspirate the media from each flask.
- Wash the cells by adding 3 mL PBS using a 5 mL pipet. Slightly tip the flask back and forth to rinse the cells then aspirate the PBS with a fresh Pasteur pipet.
- With a 2 mL pipet, add 1 mL of trypsin to each flask.
- Tip the flask in each direction to distribute the trypsin evenly then incubate the cells at 37°C for 10 minutes using a timer.
- Retrieve your flasks from the incubator and firmly tap the bottom 5 times to dislodge the cells.
- Check your cells using the microscope to ensure they are dislodged. They should appear round and move freely.
- If your cells are not detached from the flask, incubate at 37°C for an additional minute.
- When your cells are dislodged, move your flask back into the tissue culture hood and add 3 mL of PBS.
- To ensure you collect all of the cells in your flask, pipet the PBS down the bottom of the flask to wash the cells from the surface.
- Repeat this a total of 3 times.
- Note: do not take up or release all the liquid, in order to avoid bubbles.
- Transfer the suspended cells into a labeled 15 mL conical tube.
- Centrifuge your suspensions at 500 rpm for 10 min to pellet the cells.
- Carefully remove your tubes from the centrifuge and return to your bench in the main laboratory.
- Avoid agitating your tubes as the pellet is easily disrupted.
- Slowly pour the supernatent from your tubes into your 50 mL waste conical tube.
- Do not shake, tap, or flick the tubes to remove excess liquid as this will disrupt your cell pellet.
- Add 350 μL of RLT and pipet up and down to mix.
- Transfer the cell / RLT suspension to a Qiashredder column (color??).
- Be sure to label both the column insert and collection tube.
- Only 700 μL at a time can be loaded onto the column. If you have more than 700 μL, consult the teaching faculty.
- Centrifuge at 16,000 rpm for 2 min.
- Remove the columns from the collection tubes and add 700 μL of 70% ethanol, then pipet to mix.
- Transfer 700 μL of the RNA / ethanol suspension to an RNAeasy column (color??).
- Centrifuge at 8,000 rpm for 30 sec then discard the flow-through from the collection tube in your 50 mL waste conical tube.
- Repeat Steps #17 - 18 until all of the RNA / ethanol suspension has been passed through the RNAeasy column.
- Add 700 μL of RW1 and centrifuge at 8,000 rpm for 30 sec, then discard the flow-through in your 50 mL waste conical tube.
- Add 500 μL of RPE and centrifuge at 8,000 rpm for 30 sec, then discard the flow-through in your 50 mL waste conical tube.
- Add 500 μL of RPE and centrifuge at 8,000 rpm for 2 min, then discard the flow-through in your 50 mL waste conical tube.
- Move the columns into new collection tubes, then centrifuge at 16,000 rpm for 1 min.
- Move the columns into 1.5 mL tubes.
- Remove the caps from the tubes by cutting them off with scissor as they can break off in the centrifuge.
- Label the base of the tubes to ensure you keep track of your samples.
- Add 30 μL of RNase free water, then centrifuge at 8,000 rpm for 1 min.
- Alert the teaching faculty when you are done and you will be escorted to the nanodrop to check the concentrations of your purified RNA samples.
Part 2b: Generate cDNA
- Calculate the volume of your RNA solution that contains 1 μg of RNA.
- Alert the teaching faculty if this volume is greater than 9 μL.
- Add the appropriate volume of DEPC-treated water to a PCR tube.
- The total volume of RNA + DEPC-treated water should equal 9 μL.
- Add 1 μL of the oligo(dT)20 primer from the stock located at the front laboratory bench.
- Add the volume of your RNA solution that you calculated in Step #1.
- Incubate the RNA-primer mixture in the 65 °C water bath on the front laboratory bench for 5 min.
- Retrieve your tube and incubate on ice for 1 min.
- Prepare enough 'cDNA Synthesis Master Mix' for 4.5 reactions given the volumes of each component required for one reaction that are listed below. You will pipet the stock reagents at the front laboratory bench using filtered pipet tips.
- 2 μL of 10X RT buffer
- 4 μL of 25 mM MgCl2
- 2 μL of 0.1 M DTT
- 1 μL of RNaseOUT
- 1 μL of SuperScript III RT
- Add 10 μL of cDNA Synthesis Master Mix to each of you RNA-primer tubes.
- Incubate your reactions at 50 °C in the thermocycler for 50 min.
- Terminate the reactions at 85 °C in the water bath on the front laboratory bench for 5 min.
- Move your tubes to ice and incubate for 2 min.
- Add 1 μL of RNase H and incubate in the 37 °C incubator for 20 min.
- Your cDNA will be stored in the -20 °C freezer until the next laboratory session.
Part 3: Complete cell survival assay
Today you will complete the cell survival assay. As discussed previously, you will use crystal violet staining as a readout for cell abundance. While working with crystal violet is important that you wear gloves and a lab coat as it is carcinogenic and will permanently stain your clothing. It is also important that all crystal violet waste is collected in the appropriate waste containers and not poured in the sinks.
- Retrieve your 12-well plate from the 37 °C incubator in the tissue culture room and and visually inspect your cells with a microscope.
- Record your observations concerning media color, confluency, etc. in your laboratory notebook.
- Carefully carry your plate to your bench in the main laboratory.
- Aspirate the spent media from each well.
- Be sure to attach a yellow pipet tip to the Pasteur pipet on the aspirator before using.
- Change the yellow pipet between wells so as not to cross-contaminate.
- Add ~2 mL of PBS to each well and gently rock the plate back and forth to wash the cells.
- Aspirate the PBS from each well.
- Again, change the yellow pipet tip to avoid cross-contamination.
- Add 200 μL of 0.2% crystal violet dye solution to each well.
- Carefully move your plate to the rocking table and incubate for 30 min.
- Retrieve your plate and add 1 mL of dH2 to each well.
- Add the dH2 to the crystal violet by slowly pipeting it down the side of the well rather than 'blasting' it directly into the well.
- To remove the crystal violet / dH2 from your plate, flip the plate upside down over a waste container.
- Repeat Steps #8 - 9 a total of 3 times.
- If the waste dH2 from the final wash is still dark purple, consult the teaching faculty about performing additional washes.
- Add 1 mL of 1% SDS to each well and incubate your plate on your benchtop for 30 min.
- Prepare your samples for absorbance measurements using the spectrophotometer.
- Gather 12 cuvettes from the front laboratory bench and add 900 μL of 1% SDS to each.
- It may be helpful to label the 'rough' side of your cuvettes to keep track of your samples.
- Transfer the contents of each well into a cuvette.
- You may need to pipet the contents of the well to homogenize cell / dye clumps.
- To mix the liquid in your cuvette, cover the top of the cuvette with your gloved thumb and shake up and down.
- Measure and record the A570 for each of your samples, then post the data to the wiki.
- Pour the contents of your cuvettes into the crystal violet waste container and dispose of them in your benchtop waste bucket.
Reagents
Next day: Examine transcript levels in response to DNA damage
Previous day: Complete Western blot and induce DNA damage for survival and quantitative PCR assays