Difference between revisions of "20.109(S21):Module 2"
Noreen Lyell (Talk | contribs) (→Module 2) |
Noreen Lyell (Talk | contribs) (→Module 2) |
||
| Line 7: | Line 7: | ||
'''Lecturer:''' [http://be.mit.edu/directory/angela-koehler Prof. Angela Koehler]<br> | '''Lecturer:''' [http://be.mit.edu/directory/angela-koehler Prof. Angela Koehler]<br> | ||
'''Instructors:''' [http://be.mit.edu/directory/noreen-lyell Dr. Noreen Lyell], [http://be.mit.edu/directory/leslie-mcclain Dr. Leslie McClain], and [http://be.mit.edu/directory/becky-meyer Dr. Becky Meyer] <br> | '''Instructors:''' [http://be.mit.edu/directory/noreen-lyell Dr. Noreen Lyell], [http://be.mit.edu/directory/leslie-mcclain Dr. Leslie McClain], and [http://be.mit.edu/directory/becky-meyer Dr. Becky Meyer] <br> | ||
| − | '''TAs:''' <br> | + | '''TAs:''' Jeff Hsaio and Malek Kabani<br> |
==Overview== | ==Overview== | ||
Revision as of 01:27, 23 January 2021
Contents
Module 2
Lecturer: Prof. Angela Koehler
Instructors: Dr. Noreen Lyell, Dr. Leslie McClain, and Dr. Becky Meyer
TAs: Jeff Hsaio and Malek Kabani
Overview
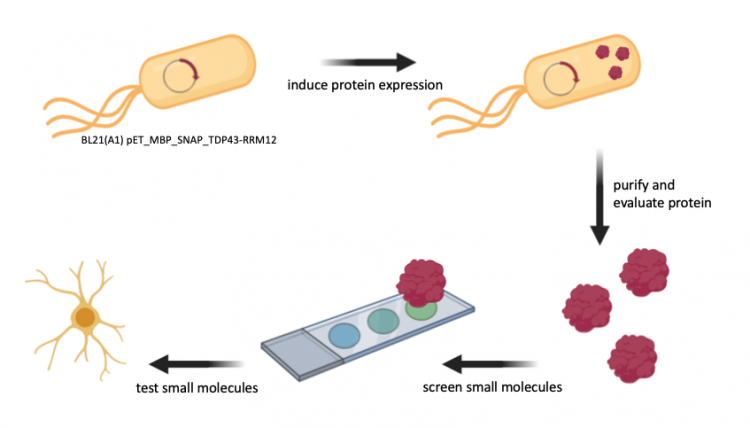
Chemical probes, or ligands, are important research tools used to explore cellular processes and therapeutic targets. The use of high-throughput and unbiased strategies to identify small molecules that bind specific biomolecules, such as proteins, can provide insight on the structure or function of targets. Additionally, a small-molecule screen can identify new chemical probes for target proteins of interest.
The small-molecule microarray (SMM) is a high-throughput method that enables the detection of protein-ligand binding. Briefly, ligands are 'printed' onto a slide and incubated with purifed protein. Unbound protein is washed from the slide and bound protein is detected using a tag on the protein of interest. Because the location of every ligand on the slide is known, the detection of protein indicates that it is bound to the ligand at that location.
In your experiment you will use SMM to identify ligands that bind to TDP43, a protein that binds both RNA and DNA. In this, TDP43 has multiple roles in transcriptional repression, pre-mRNA splicing, and translational regulation. More specifically, TDP43 is an important player in nuerodegenerative diseases.
This module has been developed thanks to the generous time and thoughtful efforts of several Koehler Laboratory members, in particular Rob Wilson.
Research goal: Identify small molecules that bind to the TDP43-RRM12 protein using small-molecule microarray screen
Lab links: day by day
M2D1: Perform protein purification protocol
M2D2: Assess purity and concentration of purified protein
M2D3: Prepare and scan small molecule microarray (SMM) slides
M2D4: Analyze SMM data to identify putative small molecule binders
M2D5: Learn best practices for mammalian cell culture
M2D6: Utilize cellular thermal shift assay (CETSA) to test putative small molecule binders
M2D7: Complete CETSA experiment and analyze data
Assignments
Journal Club presentation
Research article
References
- A method for the covalent capture and screening of diverse small molecules in a microarray format. Nature Protocols. 1:2344-2352.
- Recent discoveries and applications involving small-molecule microarrays. Chemical Biology. 18:21-28.