20.109(S18):Test protein activity using peptidyl-prolyl cis-trans isomerase assay (Day5)
Contents
Introduction
Today you will test the activity of FKBP12 using a peptidyl-prolyl isomerase (PPIase) assay. PPIases catalyze cis-trans isomerization reactions that are essential to efficient protein folding in vivo. Specifically, these enzymes isomerize peptide bonds that are N-terminal to proline residues in polypeptide chains. Without PPIases, isomerization would be the rate-limiting step in protein folding.
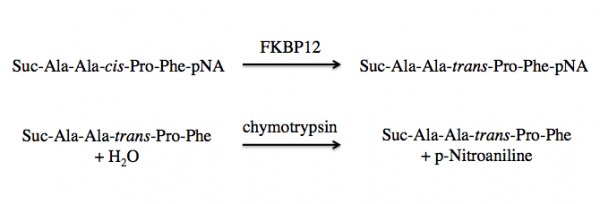
FKBP12 is a class of PPIases and the activity can be measured by quantifying the isomerization and subsequent cleavage of a substrate, suc-AAFP-pNA (also written as Suc-Ala-Ala-Pro-Phe-NA). In this method, FKBP12 catalyzes the isomeration of the cis-Ala-Pro bond to a trans-Ala-Pro bond. Then a second enzyme, chymytrypsin, cleaves the trans form of the peptide. See the reaction schematic below.
When at equilibrium in solution, approximately 88% of the commercially available suc-AAFP-pNA substrate is in the trans from, leaving only a small amount of the peptide for isomerization. Because of this, the suc-AAFP-pNA substrate is prepared in trifluoroethanol (TFE) containing lithium chloride, LiCl. The Li+ ions maintain the substrate in 60% cis form.
Upon cleavage by chymotrypsin, the release of p-nitroaniline results in a yellow color in alkaline conditions that can be measured at 405 nm. The rate of Δ405 nm in the presence versus absence of FKBP12 is used to calculate the activity.
Protocols
Part 1: Prepare samples for PPIase assay
Reagents
- FKBP12 (1 mg/mL), Abcam
- chymotrypsin, Sigma
- suc-AAFP-pNA, Sigma
Next day: Confirm ligand binding using differential scanning fluorimetry assay