Difference between revisions of "20.109(S08):Module 2"
(New page: Module 2 Day 1: Start-up protein engineering<br> (Initial reading about CaM-M13? Get calcium titration curve for WT plasmid? Need this day?...) |
MAXINE JONAS (Talk | contribs) m (36 revisions) |
||
| (35 intermediate revisions by 2 users not shown) | |||
| Line 1: | Line 1: | ||
| + | {{Template:20.109(S08)}} | ||
| − | + | <div style="padding: 10px; width: 640px; border: 5px solid #FFCC33;"> | |
| − | + | ==Module 2== | |
| − | [[ | + | '''Instructors:''' [http://web.mit.edu/be/people/jasanoff.htm Alan Jasanoff] and [http://openwetware.org/wiki/User:AgiStachowiak | Agi Stachowiak] |
| − | + | '''TA:''' [http://openwetware.org/wiki/User:Victor_S._Lelyveld |Victor Lelyveld] | |
| − | [ | + | In this experiment, you will modify a protein called inverse pericam (developed by [http://www.ncbi.nlm.nih.gov/sites/entrez?Db=pubmed&Cmd=ShowDetailView&TermToSearch=11248055&ordinalpos=5&itool=EntrezSystem2.PEntrez.Pubmed.Pubmed_ResultsPanel.Pubmed_RVDocSum Nagai et al.]) in order to change its fluorescence properties. Inverse pericam (IPC) comprises a permuted fluorescent protein linked to a calcium sensor. The ���inverse��� in the name refers to the fact that this protein shines brightly in the absence of calcium, but dimly once calcium is added. The dissociation constant <math>K_D</math> of wild-type IPC with respect to calcium is reported to be 0.2 ��M (see also figure below). Your goal will be to shift this titration curve by altering the calcium binding affinity of IPC���s calcium sensor portion. You will modify inverse pericam at the gene level using a process called site-directed mutagenesis, express the resultant protein in a bacterial host, and finally purify your mutant protein and assay its calcium-binding activity via fluorescence. In the course of this module, we will consider the benefits and drawbacks of different approaches to protein design, and the types of scientific investigations and applications enabled by fluorescently tagged biological molecules. |
| − | + | We gratefully acknowledge 20.109 instructor Natalie Kuldell for helpful discussions during the development of this module, as well as for her work in developing a [http://openwetware.org/wiki/20.109:Module_2 related module] last year. | |
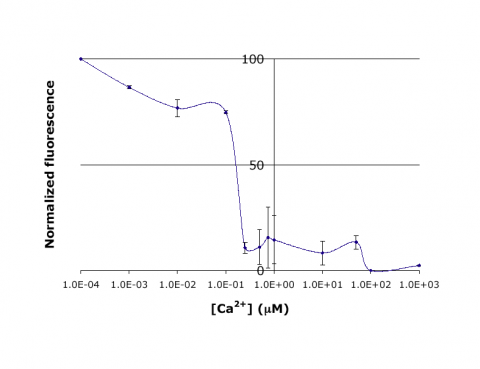
| − | [[20.109 | + | [[Image:20.109 Ca-IPC-Titr-Fig.png|thumb|center|480px|'''Titration curve for IPC.''' Shown here is sample data from the teaching lab: normalized fluorescence for wild-type inverse pericam as a function of calcium concentration. As you will later learn, an apparent <math>K_D</math> can be estimated from such a plot: it is the point on the ''x''-axis where the curve crosses ''y'' = 50%, or ~0.1 μM here.]] |
| − | ( | + | [[20.109(S08):Start-up protein engineering (Day1)| Module 2 Day 1: Start-up protein engineering]]<br> |
| + | [[20.109(S08):Site-directed mutagenesis (Day2)| Module 2 Day 2: Site-directed mutagenesis]]<br> | ||
| + | [[20.109(S08):Prepare expression system (Day3)| Module 2 Day 3: Prepare expression system]]<br> | ||
| + | [[20.109(S08):Induce protein expression (Day4)| Module 2 Day 4: Induce protein expression]]<br> | ||
| − | + | Note: week off between day 4 and day 5 of lab. | |
| − | ( | + | [[20.109(S08):Characterize protein expression (Day5)| Module 2 Day 5: Characterize protein expression]]<br> |
| + | [http://openwetware.org/wiki/20.109(S08):Assay_protein_behavior_(Day6) | Module 2 Day 6: Assay protein behavior]<br> | ||
| + | [[20.109(S08):Data analysis (Day7)| Module 2 Day 7: Data analysis]]<br> | ||
| + | [[20.109(S08):Student presentations (M2D8)| Module 2 Day 8: Student presentations]]<br> | ||
| − | [[20.109(S08): | + | [[20.109(S08): TA notes for module 2| TA notes, mod 2]] |
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
Latest revision as of 14:24, 5 June 2015
Module 2
Instructors: Alan Jasanoff and | Agi Stachowiak
TA: |Victor Lelyveld
In this experiment, you will modify a protein called inverse pericam (developed by Nagai et al.) in order to change its fluorescence properties. Inverse pericam (IPC) comprises a permuted fluorescent protein linked to a calcium sensor. The ���inverse��� in the name refers to the fact that this protein shines brightly in the absence of calcium, but dimly once calcium is added. The dissociation constant $ K_D $ of wild-type IPC with respect to calcium is reported to be 0.2 ��M (see also figure below). Your goal will be to shift this titration curve by altering the calcium binding affinity of IPC���s calcium sensor portion. You will modify inverse pericam at the gene level using a process called site-directed mutagenesis, express the resultant protein in a bacterial host, and finally purify your mutant protein and assay its calcium-binding activity via fluorescence. In the course of this module, we will consider the benefits and drawbacks of different approaches to protein design, and the types of scientific investigations and applications enabled by fluorescently tagged biological molecules.
We gratefully acknowledge 20.109 instructor Natalie Kuldell for helpful discussions during the development of this module, as well as for her work in developing a related module last year.
Module 2 Day 1: Start-up protein engineering
Module 2 Day 2: Site-directed mutagenesis
Module 2 Day 3: Prepare expression system
Module 2 Day 4: Induce protein expression
Note: week off between day 4 and day 5 of lab.
Module 2 Day 5: Characterize protein expression
| Module 2 Day 6: Assay protein behavior
Module 2 Day 7: Data analysis
Module 2 Day 8: Student presentations