Difference between revisions of "20.109(F20):M2D3"
Noreen Lyell (Talk | contribs) (→Part 1: Concentrate PF3D7_1351100 protein solution) |
Noreen Lyell (Talk | contribs) (→Part 3: Visualize PF3D7_1351100 protein using polyacrylamide gel electrophoresis (PAGE)) |
||
| Line 39: | Line 39: | ||
===Part 3: Visualize PF3D7_1351100 protein using polyacrylamide gel electrophoresis (PAGE)=== | ===Part 3: Visualize PF3D7_1351100 protein using polyacrylamide gel electrophoresis (PAGE)=== | ||
| − | + | As discussed above, SDS-PAGE is used to evaluate the purity protein. It is also possible to track the purification process using SDS-PAGE. For this, several aliquots were collected during the protein purification and concentration procedures. Each of these aliquots will be electrophoresed and the presence / absence of the protein of interest in each of the aliquots will provide information on the success of the purification. | |
| − | + | ||
<font color = #0d368e>'''To ensure the steps required for purifying the PF3D7_1351100 protein are clear, the Instructor will provide a live demonstration of this process. You should include notes regarding the procedure in your laboratory notebook!'''</font color> | <font color = #0d368e>'''To ensure the steps required for purifying the PF3D7_1351100 protein are clear, the Instructor will provide a live demonstration of this process. You should include notes regarding the procedure in your laboratory notebook!'''</font color> | ||
| − | + | #Retrieve the aliquots that were collected during the protein purification and concentration procedures. | |
| − | + | #*In addition to the aliquots that were collected from the IPTG-induced Nico(DE3) pET-28b(+)_PF3D7_1351100 cell culture, a cell lysate aliquot was collected from an uninduced cell culture. | |
| − | # | + | #To 30 μL of each aliquot, add 6 μL of Laemmli sample buffer. |
| − | #* | + | #*Prepare the cell pellet by adding 30 μL of water to a fresh microcentrifuge tube, then dip a clean pipette tip in the bacterial pellet and swirl into the water. Lastly, add 6 μL of Laemmli sample buffer. |
| − | + | #Boil all samples for 5 min in the water bath located in the chemical fume hood. | |
| − | #Boil all samples for 5 min in the water bath located in the chemical fume hood | + | #*Secure the tubes with the cap-locks located in the fume hood to ensure that the caps do not pop open during the boiling step as this will result in your sample evaporating from the tube. |
| − | #*Secure the | + | #Centrifuge samples to collect samples in the bottoms of the tubes. |
| − | # | + | |
#You will load all samples and two molecular weight standards. | #You will load all samples and two molecular weight standards. | ||
#*A pre-stained ladder will be used to track the migration of your samples through the polyacrylamide gel. | #*A pre-stained ladder will be used to track the migration of your samples through the polyacrylamide gel. | ||
#*An unstained ladder with bands of known amounts of protein will be used to estimate protein concentration in your samples. | #*An unstained ladder with bands of known amounts of protein will be used to estimate protein concentration in your samples. | ||
| − | # | + | #Load the samples and ladders into the polyacrylamid gel. |
| − | + | #Electrophorese samples at 200 V for 30-45 min. | |
| − | + | #Following electrophoresis, remove the cartridge from the electrophoresis chamber and carefully pry apart the plastic plates that encase the polyacrylamide gel. | |
| − | # | + | #With wet gloves, transfer the polyacrylamide gel to a staining box and add enough dH<sub>2</sub>O to cover the gel. |
| − | #Following electrophoresis, | + | |
| − | # | + | |
#Wash the gel for 5 min at room temperature on the rotating table. | #Wash the gel for 5 min at room temperature on the rotating table. | ||
| − | # | + | #Decant the water from the staining box in the sink. |
#*Be careful that the gel does not fall into the sink! | #*Be careful that the gel does not fall into the sink! | ||
#Repeat Steps #10-11 a total of 3 times. | #Repeat Steps #10-11 a total of 3 times. | ||
| − | #Add 50 mL of BioSafe Coomassie to the staining box and incubate | + | #Add 50 mL of BioSafe Coomassie to the staining box and incubate on the rotating table at room temperature overnight. |
#Empty the BioSafe Coomassie into the appropriate waste container in the chemical fume hood. | #Empty the BioSafe Coomassie into the appropriate waste container in the chemical fume hood. | ||
#*Be careful that the gel does not fall into the waste container! | #*Be careful that the gel does not fall into the waste container! | ||
| − | #Add 200 mL of dH<sub>2</sub>O to the staining box. | + | #Add 200 mL of dH<sub>2</sub>O to the staining box to wash the excess stain from the gel. |
| − | # | + | #Incubate on the rotating table at room temperature for 2 hrs. |
#*Replace the dH<sub>2</sub>O every 30 min. | #*Replace the dH<sub>2</sub>O every 30 min. | ||
| − | + | ||
| + | <font color = #4a9152 >'''In your laboratory notebook,'''</font color> complete the following: | ||
| + | *For each well in the polyacrylamide gel image provided by the Instructors, fully interpret the data. | ||
| + | **Do you ''expect'' to see the PF3D7_1351100 protein in the lane? Do you see PF3D7_1351100 protein? Is the band the expected size? | ||
| + | **Do you ''expect'' to see other cellular proteins in the lane? Do you see other cellular proteins? | ||
| + | *If there are unexpected results, give possible explanations. | ||
===Part 4: Measure protein concentration=== | ===Part 4: Measure protein concentration=== | ||
Revision as of 01:11, 18 July 2020
Contents
Introduction
Two metrics will be used to evaluate the success of the protein purification protocol used to express and isolate PF3D7_1351100: purity and concentration.
Protein purity
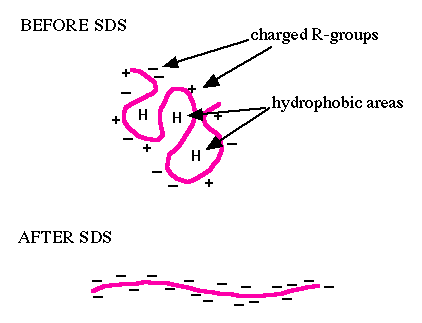
Electrophoresis is a technique that separates large molecules by size using an applied electrical field and a sieving matrix. DNA, RNA and proteins are the molecules most often studied with this technique; agarose (review M2D1!) and acrylamide gels are the two most common sieves. The molecules to be separated enter the matrix through a well at one end and are pulled through the matrix when a current is applied across it. The larger molecules get entwined in the matrix and are stalled; the smaller molecules wind through the matrix more easily and travel farther away from the well. The distance a nucleic acid or amino acid fragment travels is inversely proportional to the log of its length. Over time fragments of similar length accumulate into “bands” in the gel.
Protein concentration
To measure the concentration of your purified protein, you will use the BCA Protein Assay Reagent Kit. This kit enables colorimetric detection and quantification of the total protein within a sample. The ability to measure protein concentration is based on the detection of Cu1+ by the detection reagent, bicinchoninic acid (BCA). The Cu1+ is formed when Cu2+ is reduced by protein in an alkaline environment. Through this reduction reaction, a purple product is formed by the chelation of BCA and Cu1+ at a 2:1 ratio. This water-soluble complex has a strong absorbance at 562 nm. Because absorbance and protein concentration have a linear relationship, it is possible to compare the absorbance of an unknown protein sample to a standard curve, generated with samples of known protein concentrations, and calculate the concentration of protein in the experimental sample.
Protocols
Part 1: Concentrate PF3D7_1351100 protein solution
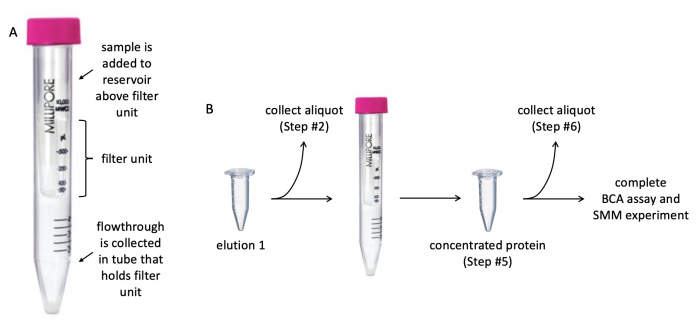
Before evaluating the purity and concentration of the PF3D7_1351100 protein, it is important to concentrate the protein solution (the protein solution is elution 1 from the previous laboratory session!). Concentrating the protein eliminates excess buffer and contaminants. To do this, a centrifugal filter with a 10 kDa cutoff will be used. The cutoff value refers to the size of the molecules that are able to pass through the filter -- molecules smaller than 10 kDa flowthrough the filter whereas molecules larger than 10 kDa are retained in the reservoir above the filter.
- Retrieve the "elution 1" sample you collected in the previous laboratory session.
- Aliquot 30 μL of the elution 1 to a fresh microcentrifuge tube.
- Label the microcentrifuge tube containing the aliquot as "elution 1" and give it to the Instructor! This aliquot will be used later when protein expression and purity are examined.
- Add the remaining contents of the original elution 1 tube to the centrifugal filter unit.
- Remove the cap from the filter unit, then pipette the purified PF3D7_1351100 protein solution into the filter unit and replace the cap.
- Centrifuge at 4500 g for 20 minutes.
- After centrifugation, transfer the liquid from the top of the filter unit to a fresh microcentrifuge tube.
- Label the microcentrifuge tube containing the liquid as "concentrated protein" and give it to the Instructor! This aliquot will be used later when protein expression and purity are examined.
- This tube contains the purified and concentrated PF3D7_1351100 protein that will be used for the SMM experiment.
- From the "concentrated protein" tube, aliquot 30 μL to a fresh microcentrifuge tube.
- Label the microcentrifuge tube containing the concentrated protein as "concentrated protein for gel" and give it to the Instructor! This aliquot will be used later when protein expression and purity are examined.
- This tube contains an aliquot of the PF3D7_1351100 protein that will be used for SDS-PAGE analysis.
Part 3: Visualize PF3D7_1351100 protein using polyacrylamide gel electrophoresis (PAGE)
As discussed above, SDS-PAGE is used to evaluate the purity protein. It is also possible to track the purification process using SDS-PAGE. For this, several aliquots were collected during the protein purification and concentration procedures. Each of these aliquots will be electrophoresed and the presence / absence of the protein of interest in each of the aliquots will provide information on the success of the purification.
To ensure the steps required for purifying the PF3D7_1351100 protein are clear, the Instructor will provide a live demonstration of this process. You should include notes regarding the procedure in your laboratory notebook!
- Retrieve the aliquots that were collected during the protein purification and concentration procedures.
- In addition to the aliquots that were collected from the IPTG-induced Nico(DE3) pET-28b(+)_PF3D7_1351100 cell culture, a cell lysate aliquot was collected from an uninduced cell culture.
- To 30 μL of each aliquot, add 6 μL of Laemmli sample buffer.
- Prepare the cell pellet by adding 30 μL of water to a fresh microcentrifuge tube, then dip a clean pipette tip in the bacterial pellet and swirl into the water. Lastly, add 6 μL of Laemmli sample buffer.
- Boil all samples for 5 min in the water bath located in the chemical fume hood.
- Secure the tubes with the cap-locks located in the fume hood to ensure that the caps do not pop open during the boiling step as this will result in your sample evaporating from the tube.
- Centrifuge samples to collect samples in the bottoms of the tubes.
- You will load all samples and two molecular weight standards.
- A pre-stained ladder will be used to track the migration of your samples through the polyacrylamide gel.
- An unstained ladder with bands of known amounts of protein will be used to estimate protein concentration in your samples.
- Load the samples and ladders into the polyacrylamid gel.
- Electrophorese samples at 200 V for 30-45 min.
- Following electrophoresis, remove the cartridge from the electrophoresis chamber and carefully pry apart the plastic plates that encase the polyacrylamide gel.
- With wet gloves, transfer the polyacrylamide gel to a staining box and add enough dH2O to cover the gel.
- Wash the gel for 5 min at room temperature on the rotating table.
- Decant the water from the staining box in the sink.
- Be careful that the gel does not fall into the sink!
- Repeat Steps #10-11 a total of 3 times.
- Add 50 mL of BioSafe Coomassie to the staining box and incubate on the rotating table at room temperature overnight.
- Empty the BioSafe Coomassie into the appropriate waste container in the chemical fume hood.
- Be careful that the gel does not fall into the waste container!
- Add 200 mL of dH2O to the staining box to wash the excess stain from the gel.
- Incubate on the rotating table at room temperature for 2 hrs.
- Replace the dH2O every 30 min.
In your laboratory notebook, complete the following:
- For each well in the polyacrylamide gel image provided by the Instructors, fully interpret the data.
- Do you expect to see the PF3D7_1351100 protein in the lane? Do you see PF3D7_1351100 protein? Is the band the expected size?
- Do you expect to see other cellular proteins in the lane? Do you see other cellular proteins?
- If there are unexpected results, give possible explanations.
Part 4: Measure protein concentration
SDS-PAGE enables you to visualize the presence of protein and provides information concerning the purity of your protein sample(s). Though comparing SDS-PAGE band intensities between samples and molecular weight standards gives an estimate of protein concentration, using a standard curve generated from samples of known protein concentration is a more precise method for measuring protein concentration.
To ensure the steps included below are clear, please watch the video tutorial (linked [[ | here]]). The steps are detailed below so you can follow along!
Prepare diluted albumin (BSA) standards
- Obtain a 100 μL aliquot of 2.0 mg/mL albumin standard stock and a conical tube of PBS from the front bench.
- Prepare your standards according to the table below using PBS as the diluent:
| Vial |
Volume of diluent (μL) | Volume (μL) and source of albumin (vial) | Final albumin concentration (μg/mL) |
|---|---|---|---|
| A | 700 | 100 of stock | 250 |
| B | 400 | 400 of A | 125 |
| C | 450 | 300 of B | 50 |
| D | 400 | 400 of C | 25 |
| E | 400 | 100 of D | 5 |
| F | 400 | 0 | 0 = blank |
Prepare Working Reagent (WR) and measuring protein concentration
- Use the following formula to calculate the volume of WR required: (# of standards + # unknowns) * 0.215 = total volume of WR (in mL).
- Prepare the calculated volume of WR by mixing 50 parts of BCA Reagent A with 1 part BCA Reagent B (50:1 or 50-fold dilution of B).
- For example, if your calculated total volume of WR is 100 mL, then mix 98 mL of A, and 2 mL of B.
- Prepare your WR in a 1.5ml microcentrifuge tube.
- Pipet 25 μL of each standard prepared in Part 4a and 25 μL of your purified TDP43-RRM12 protein into separate, wells of a flat bottom 96-well plate.
- Add 200 μL of the WR to each 25 μL aliquot of the standard OR TDP43-RRM12 protein sample (do not mix the standard and TDP43-RRM12).
- Cover your plate with plastic wrap and incubate at 37°C incubator in the back of the lab for 30 min.
- During the incubation step, check on your SDS-PAGE gel.
- Cool the plate to room temperature.
- Notify the teaching faculty you need to take your samples to the spectrophotometer plate reader.
- Obtain the absorbance at 562 nm for each well. Subtract the measurement of the Blank standard (F, after adding working reagent and being incubated) from that of all other individual standard and unknown samples.
- Generate your standard curve by plotting the Blank-corrected absorbance 562 for each albumin standard (A-F, B-F, ..., F-F) vs. its concentration in μg/mL.
- Use the standard curve to determine the protein concentration of purified TDP43-RRM12 in your sample.
Reagents
- SDS-PAGE Gels, buffer and ladder from Bio-Rad
- 4-20% polyacrylamide gels in Tris-HCl
- TGS buffer (25 mM Tris, 192 mM glycine, 0.1% (w/v) SDS, pH 8.3)
- Dual color Standard ladder and unstained ladder
- Unstained marker masses in manual linked here
- BioSafe Coomassie G-250 Stain (Bio-Rad)
- Pierce BCA protein assay (ThermoFisher)
- 6x Reducing Laemmli Sample Buffer (Boston BioProducts)
Next day: Prepare small molecule microarray (SMM) slides with purified protein