Difference between revisions of "20.109(F10): Mod 2 Day 1 Testing an engineered biological system"
(New page: {{Template:20.109(F10)}} <div style="padding: 10px; width: 640px; border: 5px solid #999999;"> =<center>Testing an engineered biological system</center>= ==Introduction== ===Why engineer b...) |
(→Reagents) |
||
| Line 10: | Line 10: | ||
If synthetic biology is to become a practical and useful engineering discipline, then it will likely need to employ engineering methodologies that characterize more mature engineering disciplines such as mechanical and electrical engineering. For example some, but not all, engineering disciplines employ well-characterized interchangeable parts. As applied to biology, interchangeable parts could mean the refinement of sequences to enable their rapid assembly (think back to how long it took us to delete 32 amino acids from the N-terminal end of GFP...a lot longer and considerably more expertise than it takes to twist a nut on a bolt!). Standardized parts might also describe families of proteins that behave in predictable ways, both individually and in combination with other protein families and sequence motifs. | If synthetic biology is to become a practical and useful engineering discipline, then it will likely need to employ engineering methodologies that characterize more mature engineering disciplines such as mechanical and electrical engineering. For example some, but not all, engineering disciplines employ well-characterized interchangeable parts. As applied to biology, interchangeable parts could mean the refinement of sequences to enable their rapid assembly (think back to how long it took us to delete 32 amino acids from the N-terminal end of GFP...a lot longer and considerably more expertise than it takes to twist a nut on a bolt!). Standardized parts might also describe families of proteins that behave in predictable ways, both individually and in combination with other protein families and sequence motifs. | ||
| − | [[Image:2CS basic.png|thumb|right| General scheme for Two Component Signaling]] Fortunately evolution appears to work with motifs and many of these motifs behave in a reasonably modular fashion. Re-used and modular motifs from nature seem sensible places to start building a library of synthetic biological parts. We'll more fully explore the idea of [http://www.partsregistry.org/Main_Page a parts registry] later in the [ | + | [[Image:2CS basic.png|thumb|right| General scheme for Two Component Signaling]] Fortunately evolution appears to work with motifs and many of these motifs behave in a reasonably modular fashion. Re-used and modular motifs from nature seem sensible places to start building a library of synthetic biological parts. We'll more fully explore the idea of [http://www.partsregistry.org/Main_Page a parts registry] later in the [http://openwetware.org/wiki/20.109(F10):_Mod_2_Day_3_Tools_for_system_engineering | module.] |
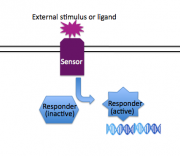
The response of bacterial cells to changing environmental conditions is one example of a modular and re-used natural theme. Hundreds of "two component signaling systems" (2CS) have been discovered in a wide range of prokaryotic cells, enabling the response of the cells to environmental changes, be they physical or chemical. Generally speaking, two component systems consist of a "sensor" protein that spans the membrane, detecting an environmental cue on its exposed face, and transducing that information to the inside of the cell through a changed activity of its cytoplasmic face. The second component of traditional two component systems is a "responder" protein, usually a transcription factor that accepts the signal from the sensor protein and consequently regulates a set of genes to provide the cell with the proper response to the external stimuli. Examples of two component signaling systems include those that regulate chemotaxis, osmoregulation, density-dependent bioluminescence, and virulence. The mechanism of two component signal transduction will be considered in depth next time. Today we'll look at how one system, the two component system that's responsive to osmolarity changes, was used to build a charismatic and novel biological system, the bacterial photography system. | The response of bacterial cells to changing environmental conditions is one example of a modular and re-used natural theme. Hundreds of "two component signaling systems" (2CS) have been discovered in a wide range of prokaryotic cells, enabling the response of the cells to environmental changes, be they physical or chemical. Generally speaking, two component systems consist of a "sensor" protein that spans the membrane, detecting an environmental cue on its exposed face, and transducing that information to the inside of the cell through a changed activity of its cytoplasmic face. The second component of traditional two component systems is a "responder" protein, usually a transcription factor that accepts the signal from the sensor protein and consequently regulates a set of genes to provide the cell with the proper response to the external stimuli. Examples of two component signaling systems include those that regulate chemotaxis, osmoregulation, density-dependent bioluminescence, and virulence. The mechanism of two component signal transduction will be considered in depth next time. Today we'll look at how one system, the two component system that's responsive to osmolarity changes, was used to build a charismatic and novel biological system, the bacterial photography system. | ||
===Bacteria as pixels=== | ===Bacteria as pixels=== | ||
| − | ''E. coli'' do not normally respond to light but [http://www.nature.com/nature/journal/v438/n7067/full/nature04405.html a recent publication ] describes a combination of genes that lead to light-responsive expression of | + | ''E. coli'' do not normally respond to light but [http://www.nature.com/nature/journal/v438/n7067/full/nature04405.html a recent publication ] describes a combination of genes that lead to light-responsive expression of ��-galactosidase in ''E. coli.'' Recall from your study of the lac operon in biology classes that ��-galactosidase is an enzyme encoded by the LacZ gene. Normally the protein cleaves the disaccharide, lactose, into two monosaccharides, galactose and glucose. |
| − | For the bacterial photography system, the "coliroid" cells producing LacZ cleave an indicator compound, giving rise to a black precipitate. When the cells are grown in the dark, transcription of LacZ is high and the indicator in the media turns black. When the cells are grown in the light, there is very little transcription of the LacZ gene and the | + | For the bacterial photography system, the "coliroid" cells producing LacZ cleave an indicator compound, giving rise to a black precipitate. When the cells are grown in the dark, transcription of LacZ is high and the indicator in the media turns black. When the cells are grown in the light, there is very little transcription of the LacZ gene and the media���s natural color (yellowish) shows through. |
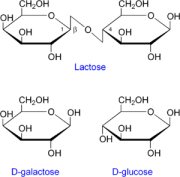
| − | [[Image:300px-Lactose etc.png|thumb|left| natural substrates and products for | + | [[Image:300px-Lactose etc.png|thumb|left| natural substrates and products for ��-galactosidase]] |
[[Image:Redskins.jpg|thumb|right| "coli-roid" example]] | [[Image:Redskins.jpg|thumb|right| "coli-roid" example]] | ||
| − | For today, you can get use to this system with a few simple tests. | + | For today, you can get use to this system with a few simple tests. You���ll plate a layer of cells on indicator medium in two dishes, and then incubate one dish in the light and the other in the dark until next time. You expect the dark-grown plate to turn a dark color and the light-grown plate to stay light. You���ll also grow some liquid cultures under the same light and dark conditions to measure ��-galactosidase activity next time. Finally, you'll familiarize yourself with the assay for ��-galactosidase activity by measuring the activity found in a bacterial strain that overproduces the enzyme. |
==Protocols== | ==Protocols== | ||
===Part 1: Test in solid media=== | ===Part 1: Test in solid media=== | ||
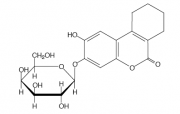
| − | [[Image:Sgal molecule.png|thumb| Molecular structure of S-gal...note its similarity to the structure of lactose (above)]] Some media has been prewarming in the 42°C waterbath for you. The media is traditional LB supplemented with ferric ammonium citrate and | + | [[Image:Sgal molecule.png|thumb| Molecular structure of S-gal...note its similarity to the structure of lactose (above)]] Some media has been prewarming in the 42°C waterbath for you. The media is traditional LB supplemented with ferric ammonium citrate and ���S-gal,��� an artificial substrate for beta-galactosidase. When the enzyme cleaves S-gal in the presence of iron, a black precipitate forms changing the color of the media. The media also has agarose so it will harden as it cools. All manipulations should be with your best sterile technique to avoid contamination of your cultures. |
<font color = purple>'''NOTE: if the TA has aliquoted the media into falcon tubes already then you can simply add antibiotics and cells and invert and pour--and thank the TA, of course!'''</font color> | <font color = purple>'''NOTE: if the TA has aliquoted the media into falcon tubes already then you can simply add antibiotics and cells and invert and pour--and thank the TA, of course!'''</font color> | ||
| Line 34: | Line 34: | ||
# Pour 15 ml of molten media into each tube and invert to mix but do not allow the media to remain at room temperature for long since it will begin to harden as it cools. | # Pour 15 ml of molten media into each tube and invert to mix but do not allow the media to remain at room temperature for long since it will begin to harden as it cools. | ||
# Add 30 μl of cells to each tube of media. Invert as before. | # Add 30 μl of cells to each tube of media. Invert as before. | ||
| − | # Pour each 15 ml volume into a Petri dishes and allow the media to harden on the bench (~30 minutes) before moving the plates into the | + | # Pour each 15 ml volume into a Petri dishes and allow the media to harden on the bench (~30 minutes) before moving the plates into the 37�� incubator that is now equipped with a red-filtered lamp. One Petri dish should be placed in a box as the ���dark��� sample and the other can be placed into the incubator media side up under the glow of the red light. |
# Expose the dishes to red filtered light (0.08-0.15W/m2 650nm range) for 24-48 hours. | # Expose the dishes to red filtered light (0.08-0.15W/m2 650nm range) for 24-48 hours. | ||
| Line 42: | Line 42: | ||
# Add 5 μl of your cells. Invert several times to mix. | # Add 5 μl of your cells. Invert several times to mix. | ||
# Split volume between 2 snap-cap tubes '''and make sure cap is snapped shut.''' You will hear a "snap" when it's really closed. If you're uncertain as to how closed your tubes are, please ask one of the faculty. | # Split volume between 2 snap-cap tubes '''and make sure cap is snapped shut.''' You will hear a "snap" when it's really closed. If you're uncertain as to how closed your tubes are, please ask one of the faculty. | ||
| − | # Place one tube in the | + | # Place one tube in the ���dark��� box and place the other in the incubator on the nutator under the red light. |
===Part 3: Beta-galactosidase assay=== | ===Part 3: Beta-galactosidase assay=== | ||
| Line 140: | Line 140: | ||
# To really really lyse the cells, vortex the tubes for 10 seconds each. You should time these precisely since you want the replicates to be treated as identically as possible. | # To really really lyse the cells, vortex the tubes for 10 seconds each. You should time these precisely since you want the replicates to be treated as identically as possible. | ||
# Start the reactions by adding 100 μl of ONPG to each tube at 10 second intervals, including your blank. Invert to mix. | # Start the reactions by adding 100 μl of ONPG to each tube at 10 second intervals, including your blank. Invert to mix. | ||
| − | # Stop the reactions by adding 250 μl of Na<sub>2</sub>CO<sub>3</sub> to each tube once sufficient yellow color has developed. | + | # Stop the reactions by adding 250 μl of Na<sub>2</sub>CO<sub>3</sub> to each tube once sufficient yellow color has developed. ���Sufficient��� is defined as yellow enough to give a reliable reading in the spectrophotometer, best between 0.3 and 1.0. It is about the amount of yellow color that you see for the yellow tips for your P200. Use today's practice assay to learn what this looks like. Try to stop the duplicate reactions at 10 second intervals and be sure to note the time you are stopping the reactions. Also be sure to remember that adding the Na<sub>2</sub>CO<sub>3</sub> makes the reactions more yellow. |
# When all your samples have been stopped, add 250 μl of Na<sub>2</sub>CO<sub>3</sub> to the blank and spin all the tubes in the microfuge for 1 minute at 13,000 RPM to pellet any cell debris. | # When all your samples have been stopped, add 250 μl of Na<sub>2</sub>CO<sub>3</sub> to the blank and spin all the tubes in the microfuge for 1 minute at 13,000 RPM to pellet any cell debris. | ||
# Move 0.7 ml of each reaction to plastic cuvettes. '''Avoid the chloroform''' that will be at the bottom of your tubes. If you add the chloroform to the cuvettes, it will "etch" the cuvette windows and mess up your readings. | # Move 0.7 ml of each reaction to plastic cuvettes. '''Avoid the chloroform''' that will be at the bottom of your tubes. If you add the chloroform to the cuvettes, it will "etch" the cuvette windows and mess up your readings. | ||
| Line 146: | Line 146: | ||
# Read the absorbance of each at 550 nm. These values reflect the amount of cell debris and differences in the plastic cuvettes themselves. | # Read the absorbance of each at 550 nm. These values reflect the amount of cell debris and differences in the plastic cuvettes themselves. | ||
# Dispose of your samples properly: liquid contents of cuvettes can go down the sink, empty cuvettes can go in the sharps containers, eppendorfs with CHCl<sub>3</sub> can go into a waste bottle in the hood. | # Dispose of your samples properly: liquid contents of cuvettes can go down the sink, empty cuvettes can go in the sharps containers, eppendorfs with CHCl<sub>3</sub> can go into a waste bottle in the hood. | ||
| − | # The | + | # The ��-gal activity in each sample is reported as "Miller Units" according to the following formula: |
<center> | <center> | ||
1 Miller Unit = <math> 1000 * \frac{(Abs{420} - (1.75*Abs{550}))}{(t * v * OD{600})}</math> | 1 Miller Unit = <math> 1000 * \frac{(Abs{420} - (1.75*Abs{550}))}{(t * v * OD{600})}</math> | ||
</center> | </center> | ||
where:<br> | where:<br> | ||
| − | '''Abs 420''' is a measure of the yellow color produced by the | + | '''Abs 420''' is a measure of the yellow color produced by the ��-gal activity.<br> |
'''Abs 550''' is a measure of cell debris. <br> | '''Abs 550''' is a measure of cell debris. <br> | ||
'''OD 600''' is a measure of the cell density.<br> | '''OD 600''' is a measure of the cell density.<br> | ||
| Line 161: | Line 161: | ||
==For Next Time== | ==For Next Time== | ||
| − | Calculate the units of the strain using the data from the | + | Calculate the units of the strain using the data from the ��-gal assays you ran, then answer the following questions: |
*Are the values for all three of the dilutions identical? | *Are the values for all three of the dilutions identical? | ||
*If so, why? If not, why not? | *If so, why? If not, why not? | ||
| Line 168: | Line 168: | ||
==Reagents== | ==Reagents== | ||
*for bacterial photography | *for bacterial photography | ||
| − | **Strain NB334 (genotype: MC4100 ara+ | + | **Strain NB334 (genotype: MC4100 ara+ ��(OmpC-lacZ) 10-25 ��envZ::KanR +pCph8 +pPL-PCBamp) |
**LB supplemented with Sgal | **LB supplemented with Sgal | ||
*** 1% Tryptone | *** 1% Tryptone | ||
| Line 179: | Line 179: | ||
** 34 ug/ul chloramphenicol | ** 34 ug/ul chloramphenicol | ||
** 25 ug/ul kanamycin | ** 25 ug/ul kanamycin | ||
| − | *for | + | *for ��-gal assay |
**Strain NB5 (genotype: BL21(DE3) +pET14b) | **Strain NB5 (genotype: BL21(DE3) +pET14b) | ||
**Z-buffer | **Z-buffer | ||
Revision as of 15:45, 23 August 2010
Testing an engineered biological system
Introduction
Why engineer biological systems
A half century of research in genetics and molecular biology has fueled remarkable progress in our scientific understanding of biological systems. As descriptions of biological systems grow more complete, it's possible to test them and use them in ever more ways. For example, an understanding of inheritance has enabled new experiments in cloning and stem cell reprogramming. An understanding of metabolism and gene expression has enabled new efforts in gene therapy and therapeutic drug production from unnatural host cells. For those who see biological science as ripe for engineering, there is an emerging discipline, synthetic biology, that hopes to enable the deliberate engineering of novel living systems. Synthetic biologists refine biology into standardized, well-characterized components to achieve defined engineering goals. If successful, the engineered biological cells, individually or as populations, may carry out molecular-scale construction projects, or exquisitely control responses to the world changing around them. In this experimental module we will consider the possibilities and limitations of biological systems as programmable materials.
Two component signaling (2CS) systems
If synthetic biology is to become a practical and useful engineering discipline, then it will likely need to employ engineering methodologies that characterize more mature engineering disciplines such as mechanical and electrical engineering. For example some, but not all, engineering disciplines employ well-characterized interchangeable parts. As applied to biology, interchangeable parts could mean the refinement of sequences to enable their rapid assembly (think back to how long it took us to delete 32 amino acids from the N-terminal end of GFP...a lot longer and considerably more expertise than it takes to twist a nut on a bolt!). Standardized parts might also describe families of proteins that behave in predictable ways, both individually and in combination with other protein families and sequence motifs.
Fortunately evolution appears to work with motifs and many of these motifs behave in a reasonably modular fashion. Re-used and modular motifs from nature seem sensible places to start building a library of synthetic biological parts. We'll more fully explore the idea of a parts registry later in the | module.The response of bacterial cells to changing environmental conditions is one example of a modular and re-used natural theme. Hundreds of "two component signaling systems" (2CS) have been discovered in a wide range of prokaryotic cells, enabling the response of the cells to environmental changes, be they physical or chemical. Generally speaking, two component systems consist of a "sensor" protein that spans the membrane, detecting an environmental cue on its exposed face, and transducing that information to the inside of the cell through a changed activity of its cytoplasmic face. The second component of traditional two component systems is a "responder" protein, usually a transcription factor that accepts the signal from the sensor protein and consequently regulates a set of genes to provide the cell with the proper response to the external stimuli. Examples of two component signaling systems include those that regulate chemotaxis, osmoregulation, density-dependent bioluminescence, and virulence. The mechanism of two component signal transduction will be considered in depth next time. Today we'll look at how one system, the two component system that's responsive to osmolarity changes, was used to build a charismatic and novel biological system, the bacterial photography system.
Bacteria as pixels
E. coli do not normally respond to light but a recent publication describes a combination of genes that lead to light-responsive expression of ��-galactosidase in E. coli. Recall from your study of the lac operon in biology classes that ��-galactosidase is an enzyme encoded by the LacZ gene. Normally the protein cleaves the disaccharide, lactose, into two monosaccharides, galactose and glucose.
For the bacterial photography system, the "coliroid" cells producing LacZ cleave an indicator compound, giving rise to a black precipitate. When the cells are grown in the dark, transcription of LacZ is high and the indicator in the media turns black. When the cells are grown in the light, there is very little transcription of the LacZ gene and the media���s natural color (yellowish) shows through.
For today, you can get use to this system with a few simple tests. You���ll plate a layer of cells on indicator medium in two dishes, and then incubate one dish in the light and the other in the dark until next time. You expect the dark-grown plate to turn a dark color and the light-grown plate to stay light. You���ll also grow some liquid cultures under the same light and dark conditions to measure ��-galactosidase activity next time. Finally, you'll familiarize yourself with the assay for ��-galactosidase activity by measuring the activity found in a bacterial strain that overproduces the enzyme.
Protocols
Part 1: Test in solid media
Some media has been prewarming in the 42°C waterbath for you. The media is traditional LB supplemented with ferric ammonium citrate and ���S-gal,��� an artificial substrate for beta-galactosidase. When the enzyme cleaves S-gal in the presence of iron, a black precipitate forms changing the color of the media. The media also has agarose so it will harden as it cools. All manipulations should be with your best sterile technique to avoid contamination of your cultures.NOTE: if the TA has aliquoted the media into falcon tubes already then you can simply add antibiotics and cells and invert and pour--and thank the TA, of course!
- Each group will need 2 sterile, empty falcon tubes (15 ml).
- Add 45 μl of antibiotics (= a cocktail you will be given that is an equal mix of 15 μl of ampicillin and 15 μl of chloramphenicol and 15 μl kanamycin) to each of the 2 tubes. The antibiotics will maintain the plasmids and reporter gene in the bacteria as the cells grow.
- Pour 15 ml of molten media into each tube and invert to mix but do not allow the media to remain at room temperature for long since it will begin to harden as it cools.
- Add 30 μl of cells to each tube of media. Invert as before.
- Pour each 15 ml volume into a Petri dishes and allow the media to harden on the bench (~30 minutes) before moving the plates into the 37�� incubator that is now equipped with a red-filtered lamp. One Petri dish should be placed in a box as the ���dark��� sample and the other can be placed into the incubator media side up under the glow of the red light.
- Expose the dishes to red filtered light (0.08-0.15W/m2 650nm range) for 24-48 hours.
Part 2: Test in liquid media
- Pipet 5 ml of LB into a 15 ml falcon tube.
- Add 15 μl of the antibiotic cocktail (= 5 μl ampicillin, 5 μl chloramphenicol, 5 μl kanamycin).
- Add 5 μl of your cells. Invert several times to mix.
- Split volume between 2 snap-cap tubes and make sure cap is snapped shut. You will hear a "snap" when it's really closed. If you're uncertain as to how closed your tubes are, please ask one of the faculty.
- Place one tube in the ���dark��� box and place the other in the incubator on the nutator under the red light.
Part 3: Beta-galactosidase assay
With this assay you will determine the amount of beta-galactosidase activity associated with a sample of bacterial cells. These cells were engineered to overproduce beta-galactosidase. You will dilute the cells and measure each dilution in duplicate to gain some confidence in the values you measure. A table is included here to help you organize your assay, but you can make one of your own if you prefer.
| Tube # | Sample | OD600 | Time started | Time stopped | Time elapsed (calculated) | OD420 | OD550 | Units (calculated) |
|---|---|---|---|---|---|---|---|---|
| 0 | blank | 0:00 | ||||||
| 1 | undiluted cells | 0:10 | ||||||
| 2 | undiluted cells | 0:20 | ||||||
| 3 | 1:10 dilution | 0:30 | ||||||
| 4 | 1:10 dilution | 0:40 | ||||||
| 5 | 1:100 dilution | 0:50 | ||||||
| 6 | 1:100 dilution | 1:00 |
- Retrieve 3 ml of the bacterial sample you'll be measuring, called "NB5."
- In eppendorf tubes, make 1 ml of a 1:10 dilution of NB5, using Zbuffer as the diluent, then use 100 ul of this dilution to make another 1:10, for a final concentration that's 1:100.
- Transfer 650 μl of the 1:10 dilution to a cuvette and measure the OD600 of this sample. Blank the spectrophotometer with Zbuffer or water. The density of the 1:10 dilution should be in the linear range of the spectrophotometer and your reading can be used to calculate the OD600 of your undiluted cells and your 1:100 dilution.
- Add 400 μl of Zbuffer to 7 eppendorf tubes labeled 0-6.
- Add 100 μl of the appropriate cell dilution to each tube. See chart above for guidance. Add 100 ul of Zbuffer to tube 0, to serve as your blank.
- Next you will lyse the cells by add 20 μl of 0.1% SDS to each eppendorf.
- To better lyse the cells, you should also add 30 μl of chloroform (CHCl3) to each tube. Do this in the hood since chloroform is volatile and toxic. You will need to hold the pipet tip close to the eppendorf as you move between the chloroform stock bottle and your eppendorfs since chloroform has a low surface tension and will drip from your pipetmen. Be sure to dispose of your pipet tips in the chloroform waste container located on the right side of the hood.
- To really really lyse the cells, vortex the tubes for 10 seconds each. You should time these precisely since you want the replicates to be treated as identically as possible.
- Start the reactions by adding 100 μl of ONPG to each tube at 10 second intervals, including your blank. Invert to mix.
- Stop the reactions by adding 250 μl of Na2CO3 to each tube once sufficient yellow color has developed. ���Sufficient��� is defined as yellow enough to give a reliable reading in the spectrophotometer, best between 0.3 and 1.0. It is about the amount of yellow color that you see for the yellow tips for your P200. Use today's practice assay to learn what this looks like. Try to stop the duplicate reactions at 10 second intervals and be sure to note the time you are stopping the reactions. Also be sure to remember that adding the Na2CO3 makes the reactions more yellow.
- When all your samples have been stopped, add 250 μl of Na2CO3 to the blank and spin all the tubes in the microfuge for 1 minute at 13,000 RPM to pellet any cell debris.
- Move 0.7 ml of each reaction to plastic cuvettes. Avoid the chloroform that will be at the bottom of your tubes. If you add the chloroform to the cuvettes, it will "etch" the cuvette windows and mess up your readings.
- Read the absorbance at 420nm. These values reflect the amount of yellow color in each tube.
- Read the absorbance of each at 550 nm. These values reflect the amount of cell debris and differences in the plastic cuvettes themselves.
- Dispose of your samples properly: liquid contents of cuvettes can go down the sink, empty cuvettes can go in the sharps containers, eppendorfs with CHCl3 can go into a waste bottle in the hood.
- The ��-gal activity in each sample is reported as "Miller Units" according to the following formula:
1 Miller Unit = $ 1000 * \frac{(Abs{420} - (1.75*Abs{550}))}{(t * v * OD{600})} $
where:
Abs 420 is a measure of the yellow color produced by the ��-gal activity.
Abs 550 is a measure of cell debris.
OD 600 is a measure of the cell density.
t is the reaction time from start to stop, measured in minutes.
v is the culture volume that you added to the reactions, measured in mls.
You can average the duplicate values for each sample unless you know of a reason not to (e.g. one tube spilled or had the incorrect amount of something added to it...)
DONE!
For Next Time
Calculate the units of the strain using the data from the ��-gal assays you ran, then answer the following questions:
- Are the values for all three of the dilutions identical?
- If so, why? If not, why not?
- What are some of the variables that affect the confidence you have in the data.
Reagents
- for bacterial photography
- Strain NB334 (genotype: MC4100 ara+ ��(OmpC-lacZ) 10-25 ��envZ::KanR +pCph8 +pPL-PCBamp)
- LB supplemented with Sgal
- 1% Tryptone
- 0.5% Yeast Extract
- 1% NaCl
- 0.03% S-gal
- 0.05% Ammonium Ferric Citrate
- 1% Low Melting Point Agarose
- 100 ug/ul ampicillin
- 34 ug/ul chloramphenicol
- 25 ug/ul kanamycin
- for ��-gal assay
- Strain NB5 (genotype: BL21(DE3) +pET14b)
- Z-buffer
- 0.06 M Na2HPO4
- 0.04 M NaH2PO4*H2O
- 0.001 M KCl
- 0.0001 M MgSO4
- 4 mg/ml ONPG
- 1M Na2CO3