Difference between revisions of "20.109(F11): Mod 2 Day 3 Tools for system engineering"
(→Part 1: Library Screen) |
(→Reagents) |
||
| Line 4: | Line 4: | ||
==Introduction== | ==Introduction== | ||
[[Image:EvolutionBiolSystem.png|thumb|left]]Biological systems are not static. Thus they can be engineered to account for changing environmental conditions, as we've seen through our examination of two component regulatory systems. In addition, they can be engineered to account for changes that occur as the cells replicate and divide over time. Indeed, evolution of biological systems away from an original specification can be viewed as a curse (it's not like computer scientists have to worry that their software programs change functions when they relaunch them!) or a blessing (evolution can find a solution we didn't ever dream of). Here we'll take the rosy view and try to harness genetic variability to improve the bacterial photography system. In particular we'll screen a library of changes in the Cph8 gene to find ones that increase the phosphatasing activity of the sensor when the cells are growing in the light. The mutants (we'll call them P+) should make the "light" color of the photographs more light by decreasing the amount of phosphorylated OmpR that stimulates LacZ transcription. | [[Image:EvolutionBiolSystem.png|thumb|left]]Biological systems are not static. Thus they can be engineered to account for changing environmental conditions, as we've seen through our examination of two component regulatory systems. In addition, they can be engineered to account for changes that occur as the cells replicate and divide over time. Indeed, evolution of biological systems away from an original specification can be viewed as a curse (it's not like computer scientists have to worry that their software programs change functions when they relaunch them!) or a blessing (evolution can find a solution we didn't ever dream of). Here we'll take the rosy view and try to harness genetic variability to improve the bacterial photography system. In particular we'll screen a library of changes in the Cph8 gene to find ones that increase the phosphatasing activity of the sensor when the cells are growing in the light. The mutants (we'll call them P+) should make the "light" color of the photographs more light by decreasing the amount of phosphorylated OmpR that stimulates LacZ transcription. | ||
| − | The region of the Cph8 protein to focus on for this purpose has been defined through traditional scientific studies of EnvZ, for example the work from Tom Silhavy's lab( [ | + | The region of the Cph8 protein to focus on for this purpose has been defined through traditional scientific studies of EnvZ, for example the work from Tom Silhavy's lab( [http://openwetware.org/wiki/PMID:_9721293 ] and [[Media:K+P- JBact98.pdf|pdf]] here). We've also been guided by the expertise of MIT's [http://web.mit.edu/biology/www/facultyareas/facresearch/laub.html Mike Laub,] whose lab studies the specificity and rewiring of two component regulatory systems. From these sources, a span of 5 contiguous amino acids can be identified as relevant for shifting the balance of EnvZ to greater kinasing or greater phosphatasing activity. These five residues in EnvZ are Alanine at amino acid 239 ("A239") through Histidine at amino acid 243 ("H243"), where mutations in the flanking residues (A239 and H243) have been shown to enhance the phosphatase activity of EnvZ and mutations in the internal residues (G240 V241 S242) enhance the kinase activity of EnvZ. The amino acid changes that modify the enzymatic activities are indicated on the figure below. Two important notes about these mutations though: First, the balance of kinase to phosphatase activities have been affected by the changes, but the mutations do not shift the reactions to fully "on" or fully "off." Second, the fusion protein of Cph1 to EnvZ, called Cph8, changes the numbering of the residues, as shown in the figure below. It's hoped, however, that the local environment of the region is similar to the natural EnvZ protein. [[Image:P+, EnvZ, Cph8 align.png]]<br> |
To complement the genetic approach for solving biological engineering puzzles, we'll also consider three more approaches in synthetic biology. The first is a [http://www.partsregistry.org/Main_Page Registry of Standard Biological Parts,] essentially a community resource that has some ready-made and useful genetic elements that can be assembled into synthetic biological devices systems. The second approach is to recapitulate the genetic network of the biological system using electronic components, making explicit some of the benefits and limitations of such an approach and the often-cited analogy between building with biology and building with computer programs or electronic components. The final approach is the one you started last time, namely the use the [http://www.tinkercell.com/ Tinkercell computer model] to simulate the behavior of the bacterial photography system, with the goal of performing "in silico" experiments that would take days or weeks to do at the bench. | To complement the genetic approach for solving biological engineering puzzles, we'll also consider three more approaches in synthetic biology. The first is a [http://www.partsregistry.org/Main_Page Registry of Standard Biological Parts,] essentially a community resource that has some ready-made and useful genetic elements that can be assembled into synthetic biological devices systems. The second approach is to recapitulate the genetic network of the biological system using electronic components, making explicit some of the benefits and limitations of such an approach and the often-cited analogy between building with biology and building with computer programs or electronic components. The final approach is the one you started last time, namely the use the [http://www.tinkercell.com/ Tinkercell computer model] to simulate the behavior of the bacterial photography system, with the goal of performing "in silico" experiments that would take days or weeks to do at the bench. | ||
==Protocols== | ==Protocols== | ||
| Line 23: | Line 23: | ||
#Hold the pulse button until you hear a beep. Listen carefully since the beep is not loud. | #Hold the pulse button until you hear a beep. Listen carefully since the beep is not loud. | ||
#'''Quickly''' remove the cuvette from the holder and '''immediately''' add the 0.5 ml volume of "SOC" media to the cells. Delaying this addition by even 1 minute has been seen to decrease transformation by 3 fold. | #'''Quickly''' remove the cuvette from the holder and '''immediately''' add the 0.5 ml volume of "SOC" media to the cells. Delaying this addition by even 1 minute has been seen to decrease transformation by 3 fold. | ||
| − | # Transfer the cells and the media back to an eppendorf tube and place the tubes on the nutator in the | + | # Transfer the cells and the media back to an eppendorf tube and place the tubes on the nutator in the 37�� incubator for 1 hour. During this incubation you can work on Parts 2, 3 and 4 of today's protocols. |
| − | # Spread 20 ul of the electroporation mix onto a Tetrazolium+Cam34+Amp25+Kan10 petri dishes. Plate 200 ul of the electroporation mix on another Tetrazolium+Cam34+Amp25+Kan10 petri dish. One of these two dilutions should have single, well-isolated colonies to examine next time. Incubate the plates at | + | # Spread 20 ul of the electroporation mix onto a Tetrazolium+Cam34+Amp25+Kan10 petri dishes. Plate 200 ul of the electroporation mix on another Tetrazolium+Cam34+Amp25+Kan10 petri dish. One of these two dilutions should have single, well-isolated colonies to examine next time. Incubate the plates at 37�� in the light until next time. |
===Part 2: Registry of Standard Biological Parts=== | ===Part 2: Registry of Standard Biological Parts=== | ||
| Line 34: | Line 34: | ||
The analogy of the DNA as computer code is not perfect. We have to set aside the presumption of an intelligent agent responsible for writing the initial program as well as accept that natural events will change the code over time (evolution leading to genetic variation--the very thing we're trying to harness in the first part of today's lab). And no good tools exist for systematically debugging the genetic code. | The analogy of the DNA as computer code is not perfect. We have to set aside the presumption of an intelligent agent responsible for writing the initial program as well as accept that natural events will change the code over time (evolution leading to genetic variation--the very thing we're trying to harness in the first part of today's lab). And no good tools exist for systematically debugging the genetic code. | ||
| − | What would make genetic code easier to write? One idea is to make it a more | + | What would make genetic code easier to write? One idea is to make it a more ���object oriented��� language, defining units of known function that could be combined in standard and predictable ways. One effort to facilitate genetic programming can be found at [http://parts.mit.edu The Registry for Standard Biological Parts]. The Registry is a catalog of parts that describe basic biological functions. For example BBa_B0010 is the part number for a transcriptional terminator. The Registry of Standard Biological Parts makes its parts freely available to interested researchers and engineers, and allows registered users to add and annotate parts. |
To familiarize yourself with the Registry, you'll design a protein-generating device for ''E. coli''. This device will consist of at least 3 parts from the Registry: an ORF, a promoter and a ribosome binding site (RBS).[[Image:SearchRegistry.png|thumb|center|550px|Registry Search Features]] | To familiarize yourself with the Registry, you'll design a protein-generating device for ''E. coli''. This device will consist of at least 3 parts from the Registry: an ORF, a promoter and a ribosome binding site (RBS).[[Image:SearchRegistry.png|thumb|center|550px|Registry Search Features]] | ||
| Line 57: | Line 57: | ||
=====Assembly of this construct===== | =====Assembly of this construct===== | ||
| − | Part of the usefulness of the Registry is that the parts conform (mostly) to a standardized assembly scheme. This scheme enables the same restriction enzymes to be used to physically piece together parts in series. The scheme does not ensure functional assemblies, however. If you have time before your cells are ready today or before the electronics exercise, then you can familiarize yourself with this [http://ginkgobioworks.com/support/ " | + | Part of the usefulness of the Registry is that the parts conform (mostly) to a standardized assembly scheme. This scheme enables the same restriction enzymes to be used to physically piece together parts in series. The scheme does not ensure functional assemblies, however. If you have time before your cells are ready today or before the electronics exercise, then you can familiarize yourself with this [http://ginkgobioworks.com/support/ "BioBrick��� assembly scheme."] |
===Part 3: Modeling biological versus electrical devices=== | ===Part 3: Modeling biological versus electrical devices=== | ||
| Line 63: | Line 63: | ||
[[Image:PhotoElectronics.jpg|thumb|left|250px|'''starting kit''']] | [[Image:PhotoElectronics.jpg|thumb|left|250px|'''starting kit''']] | ||
====Acknowledgements==== | ====Acknowledgements==== | ||
| − | Many thanks to [ | + | Many thanks to [http://openwetware.org/wiki/Tom_Knight ], [http://openwetware.org/wiki/Reshma_Shetty ] and [http://openwetware.org/wiki/Barry_Canton ] who motivated and designed an early version of this exercise. And a special shout out to Steve Wasserman for his guidance in redeveloping, troubleshooting and building this circuit to more directly reflect the experiment we're performing. Kudos also to Kelly Drinkwater, BE class of 2011 for her reworking of this material. |
===Before you Begin=== | ===Before you Begin=== | ||
| Line 226: | Line 226: | ||
==Reagents== | ==Reagents== | ||
*for electroporation | *for electroporation | ||
| − | **Strain NB462 (genotype: MC4100 ara+ | + | **Strain NB462 (genotype: MC4100 ara+ ��(OmpC-lacZ) 10-25 ��envZ::KanR +pPL-PCBamp) |
| − | **pCph8 library: see [[20.109(F11): Mod 2 Day 5 Assessing re-tuned system| Day 5]] | + | **pCph8 library (0.4 ug/ul): see [[20.109(F11): Mod 2 Day 5 Assessing re-tuned system| Day 5]] |
**SOC media | **SOC media | ||
*** 0.5% Yeast Extract | *** 0.5% Yeast Extract | ||
Revision as of 20:50, 27 October 2011
Contents
- 1 Tools for system engineering
- 1.1 Introduction
- 1.2 Protocols
- 1.2.1 Part 1: Library Screen
- 1.2.2 Part 2: Registry of Standard Biological Parts
- 1.2.3 Part 3: Modeling biological versus electrical devices
- 1.2.4 Before you Begin
- 1.2.5 Introduction to Breadboards
- 1.2.6 Part 4: CAD tool for synthetic biology
- 1.3 For Next Time
- 1.4 Reagents
Tools for system engineering
Introduction
Biological systems are not static. Thus they can be engineered to account for changing environmental conditions, as we've seen through our examination of two component regulatory systems. In addition, they can be engineered to account for changes that occur as the cells replicate and divide over time. Indeed, evolution of biological systems away from an original specification can be viewed as a curse (it's not like computer scientists have to worry that their software programs change functions when they relaunch them!) or a blessing (evolution can find a solution we didn't ever dream of). Here we'll take the rosy view and try to harness genetic variability to improve the bacterial photography system. In particular we'll screen a library of changes in the Cph8 gene to find ones that increase the phosphatasing activity of the sensor when the cells are growing in the light. The mutants (we'll call them P+) should make the "light" color of the photographs more light by decreasing the amount of phosphorylated OmpR that stimulates LacZ transcription.The region of the Cph8 protein to focus on for this purpose has been defined through traditional scientific studies of EnvZ, for example the work from Tom Silhavy's lab( [1] and pdf here). We've also been guided by the expertise of MIT's Mike Laub, whose lab studies the specificity and rewiring of two component regulatory systems. From these sources, a span of 5 contiguous amino acids can be identified as relevant for shifting the balance of EnvZ to greater kinasing or greater phosphatasing activity. These five residues in EnvZ are Alanine at amino acid 239 ("A239") through Histidine at amino acid 243 ("H243"), where mutations in the flanking residues (A239 and H243) have been shown to enhance the phosphatase activity of EnvZ and mutations in the internal residues (G240 V241 S242) enhance the kinase activity of EnvZ. The amino acid changes that modify the enzymatic activities are indicated on the figure below. Two important notes about these mutations though: First, the balance of kinase to phosphatase activities have been affected by the changes, but the mutations do not shift the reactions to fully "on" or fully "off." Second, the fusion protein of Cph1 to EnvZ, called Cph8, changes the numbering of the residues, as shown in the figure below. It's hoped, however, that the local environment of the region is similar to the natural EnvZ protein. ![]()
To complement the genetic approach for solving biological engineering puzzles, we'll also consider three more approaches in synthetic biology. The first is a Registry of Standard Biological Parts, essentially a community resource that has some ready-made and useful genetic elements that can be assembled into synthetic biological devices systems. The second approach is to recapitulate the genetic network of the biological system using electronic components, making explicit some of the benefits and limitations of such an approach and the often-cited analogy between building with biology and building with computer programs or electronic components. The final approach is the one you started last time, namely the use the Tinkercell computer model to simulate the behavior of the bacterial photography system, with the goal of performing "in silico" experiments that would take days or weeks to do at the bench.
Protocols
Before you leave today, you should examine the bacterial photograph you set up last time and document your work and your ideas about the experiment.
Part 1: Library Screen
The details for how the libraries were constructed and what kinds of changes are reasonable to expect will be considered in the next lab session. For today, you will transform a pool of DNA with degeneracies in the positions that affect the kinasing and phosphatasing activity of EnvZ. The recipient bacterial strain is identical to the bacterial photography system except that it does not harbor a plasmid encoding the light-sensing fusion protein Cph8. It does encode the OmpR-regulated LacZ gene as well as the phycobilins from a plasmid.
To carry out the transformation you will try your hand at a different technique for getting DNA into cells, namely electroporation. This method involves exposing a small but dense volume of cells to a pulse of current. The pulse momentarily opens small spaces in the lipid bilayer for the DNA to pass into the cells. As you can imagine, the cells aren't fond of such treatment, and the single most important step to help them recover is to quickly add media to the cells once they've been electroporated. The process is also very sensitive to salts in the DNA, and if you pipet too much DNA on to the cells, the extra salt may cause an electrical "arc," (you'll know this has happened from the flash of light and the "pop" you'll hear) frying the cells dead. If this happens, please get another aliquot of cells from the faculty and try the electroporation again, with less DNA. Final bit of advice: the "beep" that the electroporator makes to tell you that it's done is relatively quiet so listen carefully for it.- When you are ready to electroporate the library, retrieve an aliquot of cells from the teaching faculty, a sterile cuvette, and an aliquot of rich, pre-warmed "SOC" media.
- Put the cuvette on ice.
- Pipet 2 ul of the library DNA (library = 0.4 ug/ul) that is being held in an icebucket on the teacher's bench into your aliquot of cells.
- Let the cells and the DNA incubate on ice one minute.
- Transfer 50 ul of the cells (or more if the tube has more volume) to the chilled cuvette and recover it with the blue lid.
- Put on your safety goggles.
- Tap the cuvette on the bench so the cells rest in the bottom of the cuvette.
- With the cuvette's "nub" facing away from you, slide the cuvette into the electroporation chamber. Push the slide into the chamber until the cuvette is between the metal contacts. The lid on the cuvette will seem to block the path but in fact, it doesn't block the slider if you've lined thing up.
- Make sure the electroporator is set to "Ec2"
- Hold the pulse button until you hear a beep. Listen carefully since the beep is not loud.
- Quickly remove the cuvette from the holder and immediately add the 0.5 ml volume of "SOC" media to the cells. Delaying this addition by even 1 minute has been seen to decrease transformation by 3 fold.
- Transfer the cells and the media back to an eppendorf tube and place the tubes on the nutator in the 37�� incubator for 1 hour. During this incubation you can work on Parts 2, 3 and 4 of today's protocols.
- Spread 20 ul of the electroporation mix onto a Tetrazolium+Cam34+Amp25+Kan10 petri dishes. Plate 200 ul of the electroporation mix on another Tetrazolium+Cam34+Amp25+Kan10 petri dish. One of these two dilutions should have single, well-isolated colonies to examine next time. Incubate the plates at 37�� in the light until next time.
Part 2: Registry of Standard Biological Parts
What would it take to make DNA serve as a low-level programming language so that a genome is simply a particular program?
- DNA, like software, has an alphabet but with only four letters in the genetic code.
- Since there are proof-reading mechanisms in the "hardware," i.e. in the cell, syntax errors may be less likely to arise than in Python or Perl or C++.
- The code for cellular programs is messy but, honestly, so are computer programs. Subroutines are often dependent on one another (the cell cycle and DNA replication for example) and parts of the program get reused in useful, but complicated and unpredictable ways (seen as cross-talk in signaling pathways for example).
- Genetic code and computer code are both susceptible to viruses that highjack normally benign functions.
The analogy of the DNA as computer code is not perfect. We have to set aside the presumption of an intelligent agent responsible for writing the initial program as well as accept that natural events will change the code over time (evolution leading to genetic variation--the very thing we're trying to harness in the first part of today's lab). And no good tools exist for systematically debugging the genetic code.
What would make genetic code easier to write? One idea is to make it a more ���object oriented��� language, defining units of known function that could be combined in standard and predictable ways. One effort to facilitate genetic programming can be found at The Registry for Standard Biological Parts. The Registry is a catalog of parts that describe basic biological functions. For example BBa_B0010 is the part number for a transcriptional terminator. The Registry of Standard Biological Parts makes its parts freely available to interested researchers and engineers, and allows registered users to add and annotate parts.
To familiarize yourself with the Registry, you'll design a protein-generating device for E. coli. This device will consist of at least 3 parts from the Registry: an ORF, a promoter and a ribosome binding site (RBS).Finding a protein coding part
Try using the "catalog of parts and devices" (the leftmost icon here) to search for a part you'd like to use. Start your search in the "protein coding sequences" and then identify one of interest to you and your lab partner.
- What part number have you chosen?
- Follow the link to the data page associated with part. Is anything specified about its use or performance?
Finding a promoter part
Try finding this part with the "search parts" function (under the Registry tools list on the right-hand side of the page that's here.)
- Did you search by text, by part number, or by subpart? What worked and how well?
- What part number have you chosen?
- What can you learn about the regulation of transcription from this part's number? part's name? part's information at the Registry?
- Follow the link to the data page associated with this part. Is anything specified about its use or performance?
Finding an RBS
To find an RBS you should restrict your search to a set of the Registry's well-characterized ones from Chris Anderson's lab or the community collection, several of which are from Ron Weiss' lab.
- What part number did you choose?
- Can you anticipate the strength of this RBS in your three part construct? Why or why not?
NOW: In plain English, what behavior do you expect for your three part construct? You should include this answer in your lab notebook and if you choose to do the optional "for next time" assignment, you can use this example as a point of departure for your discussion.
Assembly of this construct
Part of the usefulness of the Registry is that the parts conform (mostly) to a standardized assembly scheme. This scheme enables the same restriction enzymes to be used to physically piece together parts in series. The scheme does not ensure functional assemblies, however. If you have time before your cells are ready today or before the electronics exercise, then you can familiarize yourself with this "BioBrick��� assembly scheme."
Part 3: Modeling biological versus electrical devices
In this section, we'll explore the issue of gain and signal strength in the context of an electrical circuit. As you work through this exercise, consider how the lessons learned from experimenting with an electronic circuit would map to the engineering of biological systems.
Acknowledgements
Many thanks to [2], [3] and [4] who motivated and designed an early version of this exercise. And a special shout out to Steve Wasserman for his guidance in redeveloping, troubleshooting and building this circuit to more directly reflect the experiment we're performing. Kudos also to Kelly Drinkwater, BE class of 2011 for her reworking of this material.
Before you Begin
Safety:
In this exercise, you'll be working with circuits connected to a battery. Although you are unlikely to seriously injure yourself, you should make a habit of unplugging / powering-off the circuit before you touch it.
Terminology:
The two terminals of the battery are labeled with a plus (high voltage) and minus (low voltage). When building a circuit, these are also referred to as power and ground, e.g. "To apply voltage across a resistor, connect one end of the resistor to power and the other end to ground."
Safety for the circuit components:
- Never directly connect the two battery terminals with wire. This is called a "short circuit" and can damage or drain your battery.
- Take note of which components require voltage to be applied in one direction and not the other. Applying voltage backwards can fry your components. Specifically:
- The OpAmp has one pin devoted to power and one devoted to ground.
- Diodes (photodiode and LED) have a positive leg and a negative leg. The positive leg must be toward power, and the negative leg toward ground. To figure out which is which, look at the body of the diode. It is round, except for one flat side, which indicates the negative leg. (To remember this, think of the flat side as looking like a minus sign.)
- Wires and resistors don't care which direction they are plugged in.
System design
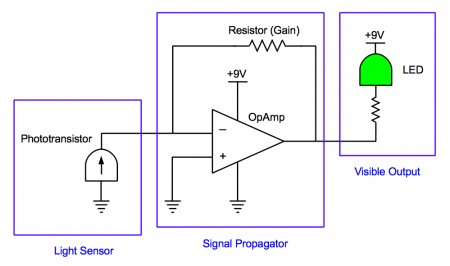
This system is a fairly simple one, consisting of only a few components. In contrast to the bacterial photography system in which the signal is propagated through protein activities, here signals are propagated as either voltage or current. When light hits the photodiode, it generates a current signal. The OpAmp takes in this current signal and produces a voltage, which signals the LED to produce light. As you can see in the schematic, the circuit contains the following parts:
- Photodiode: a light sensor. When light shines on the photodiode, its resistance decreases, and current flows through it. The photodiode is analogous to the Cph8-OmpR signaling system in the living bacterial photography system.
- OpAmp: a signal propagator. More generally, an OpAmp is a logic device that detects and amplifies a difference between the currents into its plus and minus inputs. With the addition of the feedback resistor connecting the output to the minus input, the OpAmp translates an incoming current signal into an outgoing voltage signal. The OpAmp is analogous to the transcription/translation machinery that translates an OmpR signal into synthesis of LacZ in the living bacterial photography system.
- Resistor: component which resists current flow by producing a voltage drop across it. The voltage equals the current times the resistance of the resistor; thus, the resistance sets the "gain" or amplifier strength of the system. By varying the resistance, we can vary the circuit's sensitivity to light. The resistor is analogous to the strength of the promoters and ribosome binding sites in the biological system, which raise or lower the efficiency of LacZ production in the living bacterial photography system.
- LED: a device with a detectable output (analogous to LacZ). A voltage drop across its terminals turns the green-colored light on. The small 820Ω resistor is placed in line with the LED to ensure that the current through the LED isn't too high, which can fry the LED. This small resistor has no direct analogue in the bacterial photography system. The LED is analogous to LacZ in the living bacterial photography system.
 |
 |
 |
 |
| Photodiode (sensor) | OpAmp (logic) | Resistor (gain) | LED (actuator) |
Introduction to Breadboards
(If you have used a breadboard before, feel free to skip this section.)
You will be building this circuit on a breadboard. Breadboards allow us to connect, disconnect, and move around circuit components easily and cleanly. It's much easier to build on a breadboard than to just wire everything together.
But how do those little holes connect the components together? Let's take a look at the back of a breadboard.
Each of these metal strips connects a group of holes. You can plug two wires into two holes in the same group, and those two wires will be connected. Most of the holes are grouped in short rows of five. The holes on the sides, however, are connected in long rows, forming rails that run all the way down each edge of the breadboard. These rails can be used to conveniently deliver power to many places at once.
Lighting up an LED
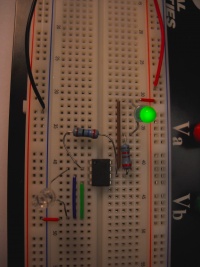
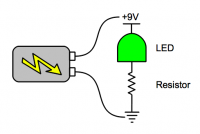
Begin with a basic circuit, to familiarize yourself with the breadboard and the LED. Examine the circuit diagram below, and then replicate it on the breadboard. Use the small (820Ω) resistor for this (stripe pattern: grey, red, black/brown, red).
 |
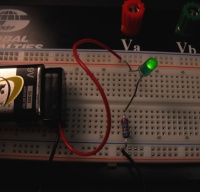 |
| Circuit diagram. | Circuit as built on breadboard. |
A few points:
- If you don't see light, check the direction of the LED and the battery, and make sure all the wires that should be connected, are connected.
- The resistor is needed to limit the current (rate of flow of electrons) through the LED. Too much current can damage it. The LEDs in your kit are strong enough that they won't burn out right away if you connect them straight across the battery, but using a resistor is still a good idea.
- The circuit you've built goes "power, LED, resistor, ground". It could also go "power, resistor, LED, ground".
Playing with the Photodiode
A photodiode is the reverse of an LED: where the LED takes an electric current and produces light, a photodiode takes in photons and produces a current. The strength of the current depends on how much light is shining on the photodiode. If you have a multimeter, you can observe this current:
- Set your multimeter to the 1mA (one milliamp) setting.
- Touch each leg of the photodiode to one of the multimeter probes.
- Shine a light on the photodiode and watch the current reading change.
Using the OpAmp
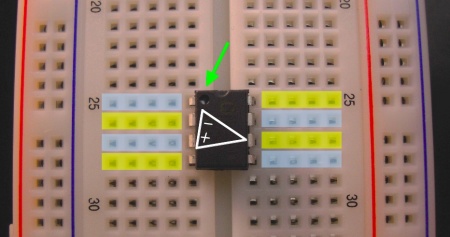
Most components have long bendable wires attached to them. The OpAmp, however, is a complicated circuit in its own right, which has been packaged inside a little black box with short pins on each side (an integrated circuit). To use the OpAmp, you must position it across the trench in the middle of the breadboard:
A few points:
- Notice how each pin gets its own row of holes. If you plugged in the OpAmp in a different position, some of the pins would be connected to each other, and the device would not work properly; all the pins need to be independent.
- Notice the small circle/divot in the upper left corner of the chip, indicated by an arrow. It lets you know which way is up. The circle/divot always marks Pin 1.
- The triangular OpAmp symbol is drawn on the back of the chip, showing which pins correspond to the OpAmp's plus input, minus input, and output (described further below). Two of the remaining pins are for power and ground.
What does an OpAmp do? OpAmp is short for "operational amplifier". Essentially, an OpAmp takes an input signal and amplifies it in various ways, depending on how other components are attached to it. OpAmps are very versatile devices. In our case, we will use a feedback resistor between the output and the minus input, which will make the OpAmp act as a current-to-voltage converter. Recall that the photodiode produces only a very small current (a few milliamps) when light shines on it. The OpAmp's job is to convert that tiny current into a much larger voltage, which will drive the LED.
Building the bacterial photography circuit
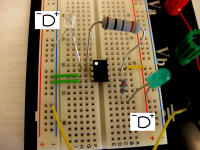
Many people find it easiest to look at this image and then try to copy it on their breadboard:
If you are not a visual person here are some written directions to help you build:
- Power the breadboard: Connect the battery leads to wires using the binding posts. Connect the power wire to the red rail at the top of the breadboard, and the ground wire to the blue rail at the bottom. (Power off the breadboard before you continue building, by disconnecting the rails or snapping the battery out of its holder.)
- Position the OpAmp across the trench in the breadboard.
- Note the small dot or half-circle marking pin 1 or the "top" of the OpAmp, as shown in the connection diagram. Power the OpAmp by connecting its voltage source inputs to the power rails (pin 7 to +9V, pin 4 to ground).
- Connect the OpAmp's plus input (pin 3) to ground.
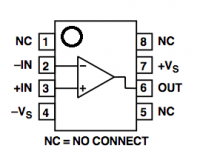 Diagram showing the connection of each pin on the OpAmp. -IN = negative input, +IN = positive input, -Vs = negative voltage supply = ground, +Vs = positive voltage supply = +9V, OUT = output, NC = pin not connected to anything. (Reproduced from data sheet.)
Diagram showing the connection of each pin on the OpAmp. -IN = negative input, +IN = positive input, -Vs = negative voltage supply = ground, +Vs = positive voltage supply = +9V, OUT = output, NC = pin not connected to anything. (Reproduced from data sheet.) - Connect the OpAmp's minus input (pin 2) to the photodiode. Remember, the photodiode is polarized. It must be connected so that the leg under the flat side is to ground.
- You'll make 2 connections to the OpAmp's output (pin 6). First, connect it to the LED and the small, 820 ohm resistor. Remember, the LED is polarized. It must be connected so that the leg under the flat side is to the resistor and the other leg is to power.
- Next, connect the OpAmp's output (pin 6) to the large, 10 mega-ohm resistor. Connect the other end of the resistor to the OpAmp's minus input (pin 2). You will change the gain of the amplifier by varying the resistance at this point (using the large resistor, a wire for zero resistance, and no wire for infinite resistance).
- Double check your connections with the circuit diagram above before you power it up.
- Test your circuit by shining a light on the photodiode and seeing if the LED responds.
Putting it all together
- Explain the role each component plays in the electronic circuit. Which part of the biological circuit does each represent?
- No doubt, building this circuit was quicker and easier than actually assembling a biological circuit -- for example, you didn't have to wait for bacteria to grow, or spend time cutting and pasting wires the way you would cut and paste DNA to make a functional plasmid. What are some traits of these components, of the breadboard, or of electronics in general, that make electrical engineering easy in this way?
- Every analogy has limitations. What are the limitations of the circuit you built as a model of the bacterial photography system? What could you do to make it more realistic?
Examining system behavior at different gain values
A circuit with very tight fully-on-or-fully-off behavior is more "digital", or switch-like, while a circuit where the LED can have a wide middle range of brightness is more "analog", or dial-like. The range of flashlight intensities that can hold the LED half-lit is a measure of the gain or strength of the amplifier. More precisely, the gain is the slope of the LED-output-vs.-photodiode-input line. Because the maximum brightness is the same for every circuit, a high-gain amplifier will cause the LED's brightness to max out even at a low level of input to the photodiode, whereas a low-gain amplifier will cause the LED's brightness to increase slowly before maxing out, as the photodiode input increases. We can tune this gain by changing the value of the gain resistor.
Initial Condition: Resistance = 10 MΩ
Right now, the OpAmp's output and minus input are connected with a 10 MΩ resistor.
- What happens to the LED when you power up the circuit?
- What happens to the LED when you shine the flashlight on the photodiode?
- Can you get the LED to hold steady at 1/2 its maximal brightness, by moving the flashlight farther away, shading it, etc?
- Sketch a graph with flashlight intensity on the x-axis and LED light intensity on the y-axis. At infinite resistance in place, is the circuit's behavior better described as a switch or a dial?
Modification: Resistance = 0Ω
Replace the 10MΩ resistor with a wire.
- What happens to the LED when you power up the circuit?
- What happens to the LED when you shine the flashlight on the photodiode?
- Can you get the LED to hold steady at 1/2 its maximal brightness?
- Add a line for this circuit to your graph. Is this circuit's behavior better described as a switch or a dial?
Alternative: Resistance = infinite Ω
Remove the wire connecting the OpAmp's output to its negative input.
- What happens to the LED when you power up the circuit?
- What happens to the LED when you shine the flashlight on the photodiode?
- Can you get the LED to hold steady at 1/2 its maximal brightness?
- Add one last line to your graph. Is this circuit's behavior better described as a switch or a dial?
Putting it all together
- How does this compare to the end of the computer simulation exercise, when you tuned the system with sliders?
- When might you want switch-like behavior in a biological system, and when might you want dial-like behavior?
- What are some everyday examples of switch-like and dial-like systems? What element or aspect plays the role of "gain" in each one?
Part 4: CAD tool for synthetic biology
If you have time and would like to revisit Tinkercell, you can run additional simulations or gather more information about the system you're experimenting with.
DONE!
For Next Time
1. Draft your Materials and Methods section for your Mod 2 research article. You can describe the strains and plasmids you've used, the mechanics of taking a bacterial photograph, the b-gal assay and the library screen as far as through the electroporation. The due date for this assignment will vary depending on if you are presenting a Journal Club article next time as well.
- If you ARE NOT giving a journal club talk next time, then this draft is due next time
- If you ARE giving a journal club talk next time, then this draft is due before lab, one week from today.
2. If you are giving a journal club talk, the slides for your presentation should be uploaded to the Stellar website that is associated with our class. The presentation order will be determined by the order that your finished slides are uploaded.
3. (OPTIONAL): Comparisons of biological engineering and electrical engineering. Today's lab work may have made clear how much easier it is to assemble electrical circuits and run computer simulations of them than it is to assemble and test biological circuits. For example, it takes seconds to swap in a new resistor into your circuit or modulate an experimental parameter in the CAD tool, but it takes a few days to assemble DNA fragments (BioBricks or not). What other comparisons can you make that emphasize the
- sensitivity of system performance
- resolution
- fabrication
- repair
- ???
of the two types of circuits.
Reagents
- for electroporation
- Strain NB462 (genotype: MC4100 ara+ ��(OmpC-lacZ) 10-25 ��envZ::KanR +pPL-PCBamp)
- pCph8 library (0.4 ug/ul): see Day 5
- SOC media
- 0.5% Yeast Extract
- 2% Tryptone
- 10 mM NaCl
- 2.5 mM KCl
- 10 mM MgCl2
- 10 mM MgSO4
- 20 mM Glucose
- Tetrazolium(lac) + Amp + Cam plates (per liter)
- 25.5 g Antibiotic Medium #2 ("Difco" brand)
- 50 mg Tetrazolium indicator/L
- 1 % lactose
- 25 ug/ml ampicillin
- 34 ug/ml chloramphenicol
- 10 ug/ml kanamycin
| Part | Source | Catalog # | ~Cost |
| Breadboard | Global (UDoIt) | PB83E | 17 |
| Alligator clips | Mueller Electric Co (DigiKey) | 314-1010-ND | 0.5 |
| 9V snap battery cover | Philmore (UDoIt) | BC120 | 1.5 |
| 9V battery | Energizer (UDoIt) | EN22, industrial | 1.6 |
| Flashlight | LED keychain lights (Walgreen's) | 693315-805697 | 2 |
| LED | AllElectronics | LED-2, 5mm green T1 3/4 | 0.5 |
| Resistor, 820 ohm | NTE (UDoIt) | QW182, 820 ohm, 2% | 0.5 |
| OpAmp | Analog Devices Inc (DigiKey) | AD8031ANZ | 3 |
| Resistor, 10 Mohm | NTE (UDoIt) | 2W610 | 1 |
| Photodiode | Leditech (AllElectronics) | LT959X-91-0125 | 0.5 |
| Wires | Global (UDoIt) | WK-3 | 3 |
| Resistor color code card | Elenco (UDoIt) | CC-100 | 1 |
| Clear case to hold parts | Flambeau (UDoIt) | DB221 | 2 |