Difference between revisions of "20.109(S07): Start-up expression engineering"
| Line 131: | Line 131: | ||
Everyone's starting strain is called FY2068, which has the following genotype:<center><br>MAT(alpha) <i>ura3-52 his3D200 leu2D1 lys2-128delta </i> <br></center>This genotype tells you that the strain is haploid and of the "alpha" mating type. The genotype also tells you that the strain cannot make its own uracil, histidine, leucine or lysine due to the indicated mutations in the <i> URA3, HIS3, LEU2 </i> and <i> LYS2</i> genes. These mutations have insignificant effects when the strain is grown on rich or "complete" media but no growth occurs when one of those needed media components is left out, or "dropped out" as such media is usually described. | Everyone's starting strain is called FY2068, which has the following genotype:<center><br>MAT(alpha) <i>ura3-52 his3D200 leu2D1 lys2-128delta </i> <br></center>This genotype tells you that the strain is haploid and of the "alpha" mating type. The genotype also tells you that the strain cannot make its own uracil, histidine, leucine or lysine due to the indicated mutations in the <i> URA3, HIS3, LEU2 </i> and <i> LYS2</i> genes. These mutations have insignificant effects when the strain is grown on rich or "complete" media but no growth occurs when one of those needed media components is left out, or "dropped out" as such media is usually described. | ||
| − | Everyone will delete a SAGA subunit of their choosing by replacing it with a selectable marker, namely the <i> URA3 </i> gene. Successful replacement of the SAGA subunit with the <i> URA3 </i> gene will restore growth of the strain on media lacking uracil ("SC" for "synthetic complete" "minus ura" for the absence of uracil). | + | Everyone will delete a SAGA subunit of their choosing by replacing it with a selectable marker, namely the <i> URA3 </i> gene. Successful replacement of the SAGA subunit with the <i> URA3 </i> gene will restore growth of the strain on "SC-ura," which is media lacking uracil ("SC" for "synthetic complete;" "minus ura" for the absence of uracil). The primers you design today will have sequences identical to the URA3 gene, shown below. The homology will enable the primers to anneal to the <i> URA3 </i> gene and amplify it during the polymerase chain reactions you will perform at the end of lab today. These reactions will be more fully explained later in the experimental module. |
Everyone's primers will also include "tails" that will later allow the amplified <i> URA3 </i> gene to replace the SAGA subunit gene of interest. These tails must be at least 39 bases long to allow for sufficient specificity and frequency of recombination once transformed for recombination into the yeast cell. This allows 20 bases for annealing to the <i>URA3 </i> gene since most oligonucleotide synthesis companies change their pricing structure and recommendations for oligos longer than 59 bases. Limitations in the synthesis technology impose the 59 base limit, but it turns out to be minimally intrusive for experiments like these. | Everyone's primers will also include "tails" that will later allow the amplified <i> URA3 </i> gene to replace the SAGA subunit gene of interest. These tails must be at least 39 bases long to allow for sufficient specificity and frequency of recombination once transformed for recombination into the yeast cell. This allows 20 bases for annealing to the <i>URA3 </i> gene since most oligonucleotide synthesis companies change their pricing structure and recommendations for oligos longer than 59 bases. Limitations in the synthesis technology impose the 59 base limit, but it turns out to be minimally intrusive for experiments like these. | ||
====Designing the "forward" primer==== | ====Designing the "forward" primer==== | ||
| + | #You and your partner should begin by opening a new MSWord document to create a primer record for the sequences you are designing today. Put your names at the top, your team color, today's date and a short description of what you are trying to do. | ||
| + | #Begin your primer design by retreiving the genomic DNA sequence for the URA3 gene as it's listed at [http://www.yeastgenome.org/ SGD]. You will need 20 bases from the start of the gene to serve as the "landing sequence" for this primer. Copy these 20 bases to your MSWord document. | ||
| + | #It is extremely important to know the melting temperature for this 20 base landing sequence. The first few rounds of PCR must be performed at a temperature that allows these 20 bases to bind. The "tails" of the primer will still be present during these first rounds of PCR but will have no complementary sequences to which they can anneal. The melting temperature depends on both the length of the landing sequence, its GC content, and any secondary structures that the primer can adopt. A well-designed primer will have short hairpins (if any), its melting temperature will be around 60°C, and if possible its GC content will be about 50%. There are several websites to help you evaluate these aspects of your primer. Try to copy the 20 bases of landing sequence into the [http://www.cybergene.se/primer.html Cybergene website]. Note on your MSWord document the Tm, | ||
| + | |||
| + | |||
| + | |||
====Designing the "reverse primer==== | ====Designing the "reverse primer==== | ||
===Part 3: PCR=== | ===Part 3: PCR=== | ||
Revision as of 16:55, 30 December 2006
Contents
Introduction
From your work so far this term, you have a good understanding of (at least) two fundamental concepts. From our first experimental module it should be clear that the genetic program for running a cell is readable (through sequencing), writable (through molecular biological techniques and synthesis) and somewhat, though not perfectly understandable. Recall how a genetic part can be as small as 13 base pairs, BBa_B0032 for example, to help carry out the work of protein production by enabling a huge protein/RNA complex, the ribosome, to bind an RNA molecule. From our second experimental module, it should be clear that a cell's programming doesn't end at protein production but rather that proteins are dynamic (chemically and spatially). They react to changes in the environment with great speed and sensitivity. We saw that proteins could be considered "digital" information, either present or absent, but are perhaps better thought of as "tunable" analog data since, as far as the cell is concerned, proteins are relevant only when functional, and function can be governed by a regulating protein stability, localization and modifications. Thus, from the work we've done so far this term, you may have the idea that gene expression in a cell is powered by the central dogma (DNA making RNA making protein) and then modulated by varying the properties of proteins once they are made. This strategy minimizes reaction times but is, energetically speaking, quite wasteful. Why make a protein if it's not useful? In this experimental module we'll see how nature has refined the genetic programming language so a cell's proteins are not all constitutively expressed but rather are finely regulated at the level of transcription initiation. This turns out to be only one of many points for regulation but it's an important one.
In the upcoming experiment we'll also see how nature has overcome another design issue, namely space constraints. A cell's dimensions do not increase linearly with DNA content. Instead, eukaryotic cells remain compartmentalized and compact the DNA by wrapping it around assemblies of histone proteins called nucleosomes. Nucleosomes wrap around eachother to form chromatin. This solves the space issue, allowing a meter or so of DNA to be crammed into a space perhaps 10 um across, but creates a new problem. Wrapped DNA is less accessible to the transcription and replication machinery. Gene expression becomes newly and intimately related to chromatin dynamics. This new problem is overcome by other multiprotein complexes that interact with the DNA-wrapping proteins. Nucleosomes are redistributed around genes that are "active" though it remains unclear if this redistribution is a cause or a consequence of the activity. We'll study one chromatin-remodeling complex called SAGA in this experimental module.
The name "SAGA" is an acronym for "Spt-Ada-Gcn5-acetyltransferase." Before describing each of these components, it's important to note that biochemically similar complexes are found in many (all?) of the eukaryotic cells that have been studied. Even more remarkable, these SAGA complexes have similar (identical?) numbers of protein subunits and the proteins have noteable sequence homologies, suggesting conserved functions even in cells with diverse life-styles like yeast and human cells. Natural processes like development and division and disease states like cancer can be understood at the level of transcription (mis)regulation. Chromatin remodeling is required for appropriate gene expression which is, in turn, required for healthy cell behaviors. Thus, there is good reason to believe that an understanding of how SAGA works in yeast can give us insight into its role in cells more medically relevant, like human.
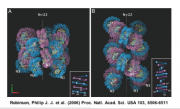
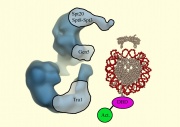
A combination of biochemical and genetic data initially suggested that an enzymatic activity, namely a histone-acetyl transferase, encoded by the GCN5 gene in S. cerevisiae existed as a large protein complex called that the authors named "SAGA" [Grant et al 1997]. Nineteen proteins, including GCN5, associate to form SAGA, though surprisingly not all the proteins are absolutely required for the cell to live. Delete a nonessential subunit and the cell can still grow and divide, although sometimes with impaired functions. A table of all 19 SAGA subunits, including information about their essential or non-essential nature, is included as part of today's protocol. A recent structure for the complex was elucidated through electron microscopy [Wu et al 2004]. The SAGA structure, as shown above, can be imagined to "dock" with the DNA and associated proteins, allowing us to imagine some very elegant models for how chromatin-modifications might be performed and regulated. Many terrific and exciting experiments can be designed to test these models. Your experiment will be a systematic examination of the non-essential SAGA subunits and their role in gene expression. Today you will design some primers to delete from the yeast genome a nonessential subunit of your choosing. Later in this experimental module, you will examine your mutated strain for changes in gene expression, looking for new phenotypes associated with the SAGA-subunit deletion as well as looking by DNA microarray for genes whose expression is affected by the loss of this subunit.
Protocols
Part 1: Choosing a SAGA subunit
You should begin by learning a little about the different classes of SAGA subunits. What does "Spt" stand for (and how do you pronounce that?!)? What is a TAF? Why is Ada5 also called Spt20?
The nomenclature for S. cerevisiae is precise and helpful. Wild type genes are normally given an italicized, three letter acronym based on the phenotype of a mutation in that gene. So a HIS gene is unable to make histidine if the gene is defective (of course, dead cells all have only one phenotype so this presumes loss of the gene product doesn't kill the cell...). Since there exist several genes that can give rise to similar phenotypes, associated genes are given a number, e.g. HIS3, HIS4,, etc. To describe recessive mutant alleles, lower case letters are used. So a strain that is his3 has a mutation that affects the function of the HIS3 gene. Since there can be several different mutations described for any given gene, a second number gets associated with the mutant, e.g. his3-1 or his3delta200. Proteins are distinguished from DNA by capitalizing only the first letter of the gene product: the HIS3 gene makes the His3 protein. Naturally there are exceptions to these rules, but in general you can pretty confidently follow them.
Begin aquainting yourself with the SAGA subunits by copying the following tables to your wiki userpage, then finding relevant information in the Saccharomyces Genome Database for the subunits. You can also search Pubmed to find out a little more about the role of the subunits in gene expression. Do not shortchange yourself on this part of the experiment, since you will be working with the subunit you choose today for the rest of the module.
SAGA subunits, S. cerevisiae
| Ada subunits | size,chromosome,null p-type | notes |
|---|---|---|
| Ada1 (aka HFI1, SUP110, SRM12, GAN1) | 1.467 kb/489 aa, Chr. XVI, viable |
|
| Ada2 (aka SWI8) | 1.305 kb/434aa, Chr. IV, viable |
|
| Ada3(aka NGG1, SWI7) | 2.109 kb/702aa, Chr. IV, viable |
|
| Gcn5 (aka ADA4, SWI9) | 1.32 kb/439aa, Chr. VII, viable |
|
| Ada5 (aka SPT20) | 1.815 kb/604aa, Chr. XV, viable |
| Spt subunits | size, chromosome, null p-type | notes |
|---|---|---|
| Spt3 | 1.014 kb/337aa, Chr. IV, viable |
|
| Spt7(aka GIT2) | 3.999 kb/1332aa, Chr. II, viable |
|
| Spt8 | 1.809 kb/602aa, Chr. XII, viable |
|
| Spt20 (aka Ada5) | 1.815 kb/604aa, Chr. XV, viable |
| TAF subunits | size, chromosome, null p-type | notes |
|---|---|---|
| TAF5 (aka TAF90) | 2.397 kb/798aa, Chr. II, inviable | |
| TAF6 (aka TAF60) | 1.551 kb/516aa, Chr. VII, inviable | |
| TAF9 (aka TAF17) | 0.474 kb/157aa, Chr. XIII, inviable | |
| TAF10 (aka TAF23, TAF25) | 0.621 kb/206aa, Chr. IV, inviable | |
| TAF12(aka TAF61, TAF68) | 1.620 kb/539aa, Chr. IV, inviable |
| Tra1 subunit | size, chromosome, null p-type | notes |
|---|---|---|
| Tra1 | 11.235 kb/3744aa, Chr. VIII, inviable |
| other subunits | size, chromosome, null p-type | notes |
|---|---|---|
| Sgf73 | 1.974 kb/657aa, Chr. VII , viable |
|
| Sgf29 | 0.779 kb/259aa, Chr. III, viable |
|
| Sgf11 | 0.3 kb/99aa, Chr.XVI, viable |
|
| Ubp8 | 1.416 kb/471aa, Chr. XIII, viable |
|
| Sus1 | gene with intron, Chr. II, viable |
Part 2: Designing deletion oligos
Everyone's starting strain is called FY2068, which has the following genotype:MAT(alpha) ura3-52 his3D200 leu2D1 lys2-128delta
Everyone will delete a SAGA subunit of their choosing by replacing it with a selectable marker, namely the URA3 gene. Successful replacement of the SAGA subunit with the URA3 gene will restore growth of the strain on "SC-ura," which is media lacking uracil ("SC" for "synthetic complete;" "minus ura" for the absence of uracil). The primers you design today will have sequences identical to the URA3 gene, shown below. The homology will enable the primers to anneal to the URA3 gene and amplify it during the polymerase chain reactions you will perform at the end of lab today. These reactions will be more fully explained later in the experimental module.
Everyone's primers will also include "tails" that will later allow the amplified URA3 gene to replace the SAGA subunit gene of interest. These tails must be at least 39 bases long to allow for sufficient specificity and frequency of recombination once transformed for recombination into the yeast cell. This allows 20 bases for annealing to the URA3 gene since most oligonucleotide synthesis companies change their pricing structure and recommendations for oligos longer than 59 bases. Limitations in the synthesis technology impose the 59 base limit, but it turns out to be minimally intrusive for experiments like these.
Designing the "forward" primer
- You and your partner should begin by opening a new MSWord document to create a primer record for the sequences you are designing today. Put your names at the top, your team color, today's date and a short description of what you are trying to do.
- Begin your primer design by retreiving the genomic DNA sequence for the URA3 gene as it's listed at SGD. You will need 20 bases from the start of the gene to serve as the "landing sequence" for this primer. Copy these 20 bases to your MSWord document.
- It is extremely important to know the melting temperature for this 20 base landing sequence. The first few rounds of PCR must be performed at a temperature that allows these 20 bases to bind. The "tails" of the primer will still be present during these first rounds of PCR but will have no complementary sequences to which they can anneal. The melting temperature depends on both the length of the landing sequence, its GC content, and any secondary structures that the primer can adopt. A well-designed primer will have short hairpins (if any), its melting temperature will be around 60°C, and if possible its GC content will be about 50%. There are several websites to help you evaluate these aspects of your primer. Try to copy the 20 bases of landing sequence into the Cybergene website. Note on your MSWord document the Tm,
Designing the "reverse primer
Part 3: PCR
- pRS406 template
DONE!