Difference between revisions of "20.109(F20):M3D3"
Noreen Lyell (Talk | contribs) (→Introduction) |
Noreen Lyell (Talk | contribs) (→Protocols) |
||
| Line 17: | Line 17: | ||
==Protocols== | ==Protocols== | ||
| − | ===Part 1: Analyze ethanol yield data=== | + | ===Part 1: Review experimental setup=== |
| + | |||
| + | |||
| + | ===Part 2: Analyze ethanol yield data=== | ||
==Reagents list== | ==Reagents list== | ||
Revision as of 01:44, 20 July 2020
Contents
Introduction
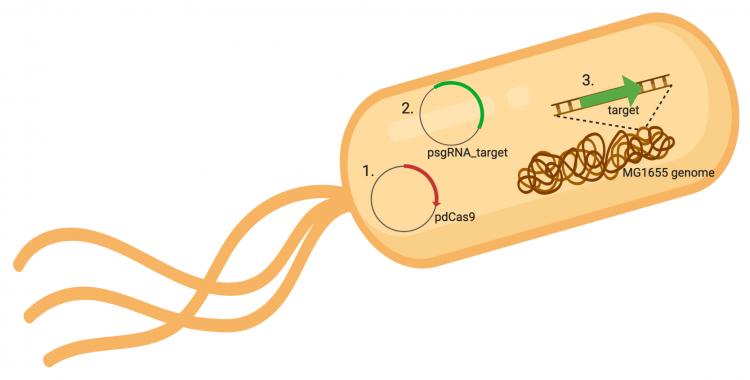
The CRISPRi system involves three genetic elements: the target gene within the host genome, the pdCas9 plasmid, and the pgRNA_target plasmid. Though the target genome is native to the host cell, the plasmids must be transformed into the cell and maintained using antibiotic selection. Throughout this module, we have learned about and worked with the CRISPRi plasmids as individual units, but now we will consider the system as a whole in the context of gene regulation.
In the previous laboratory session, you co-transformed pdCas9 and your pgRNA_target plasmids into E. coli MG1655. The colonies present on your LB plates containing chloramphenicol and ampicillin should carry both plasmids in that they were able to survive selection by both antibiotics. The promoter (pJ23119) driving expression of your gRNA sequence in the gRNA_target plasmid is constitutively active. This means that RNAP is not prohibiting from binding and transcription of the gRNA target sequence is therefore not inhibited. Therefore, your gRNA_target is always present in the MG1655 cells and binding to the target sequence within the genome.
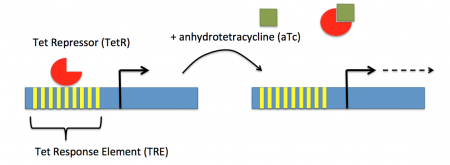
In contrast, expression of the gene encoding Cas9 within pdCas9 is regulated by an inducible promoter (pLtetO-1). An inducible promoter is 'off' unless the appropriate molecule is present to relieve repression. In the case of our system, expression of the gene encoding Cas9 is inhibited due to the use of a tet-based promoter construct. Tet is shorthand for tetracycline, which is an antibiotic that inhibits protein synthesis through preventing the association between charged aminoacyl-tRNA molecules and the A site of ribosomes. Bacterial cells that carry the tet resistance cassette are able to survive exposure to tetracycline by expressing genes that encode an efflux pump that 'flushes' the antibiotic from the bacterial cell. To conserve energy, the tet system is only expressed in the presence of tetracyline. In the absence of tetracycline, a transcription repressor protein (TetR) is bound to the promoter upstream of the tet resistance cassette genes. When tetracycline is present, the molecule binds to TetR causing a confirmational resulting in TetR 'falling off' of the promoter. In the CRISPRi system, the tet-based promoter construct upstream of the gene that encodes Cas9 is 'off' unless anhydrotetracyline (aTc), an analog of tetracyline, is added to the culture media.Taken together, the gRNA-target molecule is constitutively transcribed and, as stated above, always present / binding to the target. The dCas9 protein is only present when aTc is added. Thus, gene expression is only altered when aTc is present. As represented by the schematic below, the gRNA_target 'seeks out' the target within the host genome and recruits dCas9 to the site. When associated with the target / gRNA_target complex, dCas9 binds to the site and acts as a 'roadblock' by prohibiting RNAP access to the sequence. Because the gene of interest is not able to be transcribed, the protein encoded by that gene is not synthesized. In our experiments, we hypothesize that the absence of specific proteins, or enzymes, in the fermentation pathway will increase the yield of either ethanol or acetate.
Protocols
Part 1: Review experimental setup
Part 2: Analyze ethanol yield data
Reagents list
Next day: Examine genomic features in gRNA-targeted sequences